Bas Groot Koerkamp
for the Alzheimers Disease Neuroimaging Initiative
Reproducible radiomics through automated machine learning validated on twelve clinical applications
Aug 19, 2021
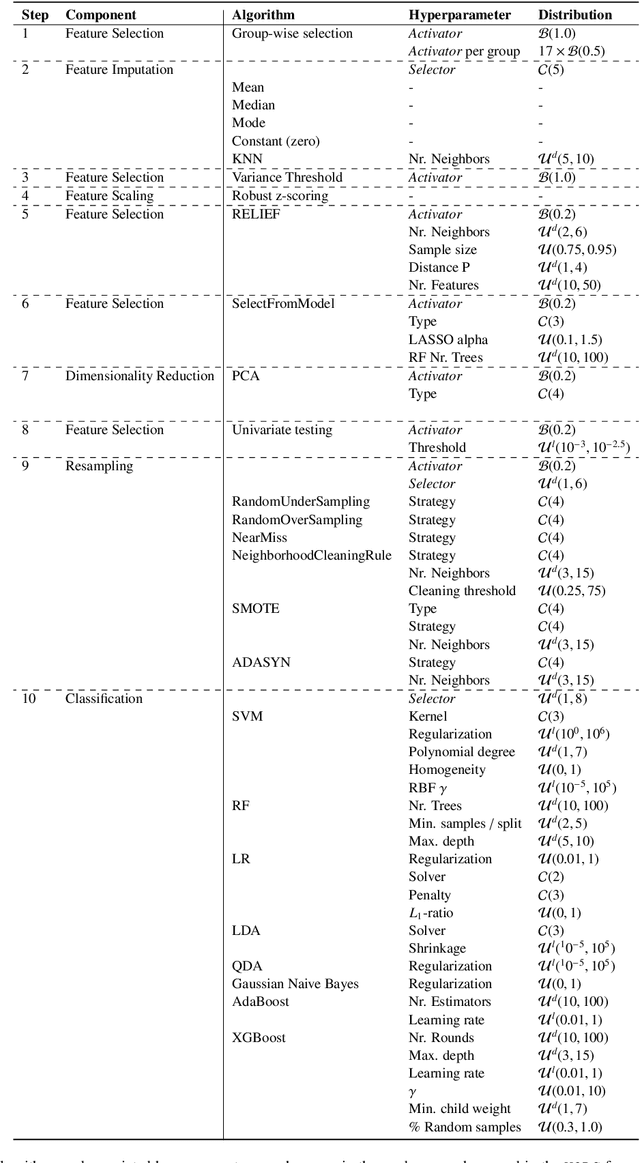
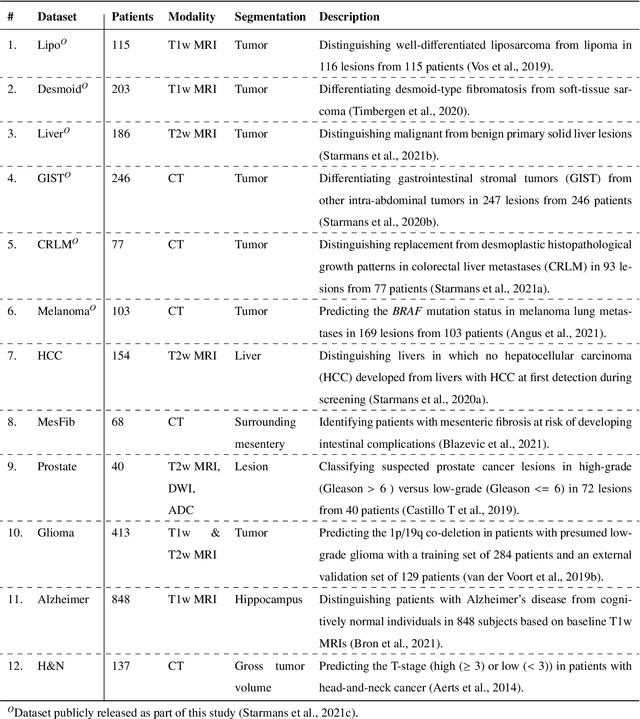
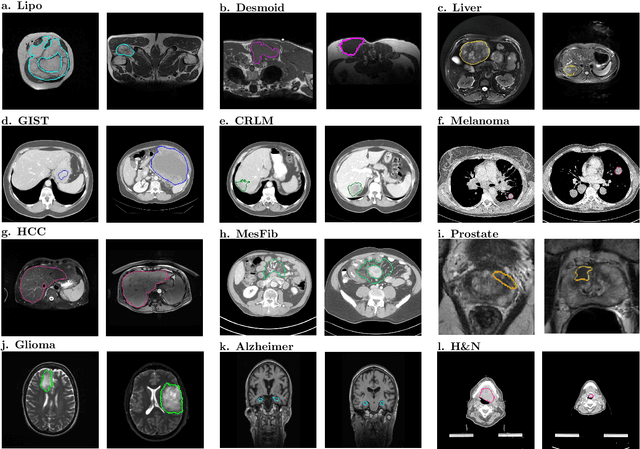
Abstract:Radiomics uses quantitative medical imaging features to predict clinical outcomes. While many radiomics methods have been described in the literature, these are generally designed for a single application. The aim of this study is to generalize radiomics across applications by proposing a framework to automatically construct and optimize the radiomics workflow per application. To this end, we formulate radiomics as a modular workflow, consisting of several components: image and segmentation preprocessing, feature extraction, feature and sample preprocessing, and machine learning. For each component, a collection of common algorithms is included. To optimize the workflow per application, we employ automated machine learning using a random search and ensembling. We evaluate our method in twelve different clinical applications, resulting in the following area under the curves: 1) liposarcoma (0.83); 2) desmoid-type fibromatosis (0.82); 3) primary liver tumors (0.81); 4) gastrointestinal stromal tumors (0.77); 5) colorectal liver metastases (0.68); 6) melanoma metastases (0.51); 7) hepatocellular carcinoma (0.75); 8) mesenteric fibrosis (0.81); 9) prostate cancer (0.72); 10) glioma (0.70); 11) Alzheimer's disease (0.87); and 12) head and neck cancer (0.84). Concluding, our method fully automatically constructs and optimizes the radiomics workflow, thereby streamlining the search for radiomics biomarkers in new applications. To facilitate reproducibility and future research, we publicly release six datasets, the software implementation of our framework (open-source), and the code to reproduce this study.
Towards Unsupervised Cancer Subtyping: Predicting Prognosis Using A Histologic Visual Dictionary
Mar 12, 2019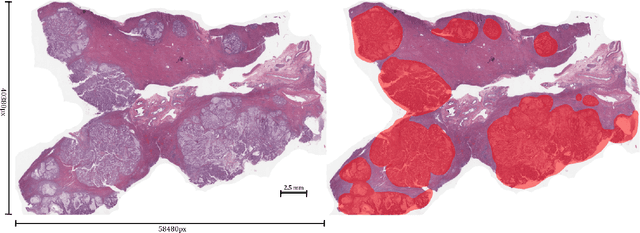
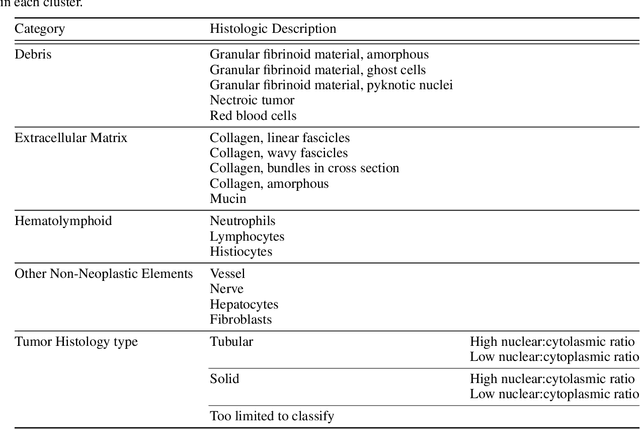
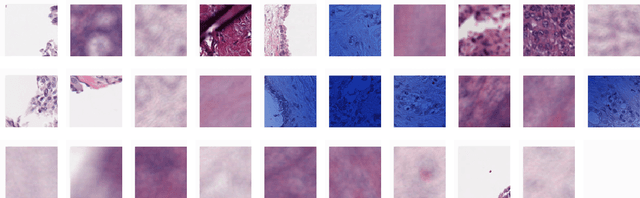
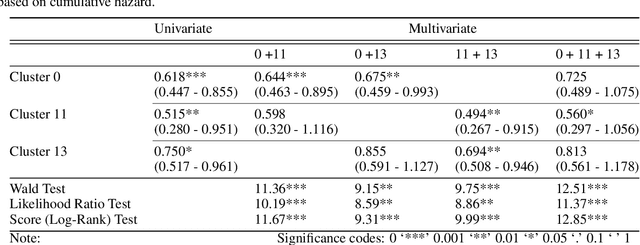
Abstract:Unlike common cancers, such as those of the prostate and breast, tumor grading in rare cancers is difficult and largely undefined because of small sample sizes, the sheer volume of time needed to undertake on such a task, and the inherent difficulty of extracting human-observed patterns. One of the most challenging examples is intrahepatic cholangiocarcinoma (ICC), a primary liver cancer arising from the biliary system, for which there is well-recognized tumor heterogeneity and no grading paradigm or prognostic biomarkers. In this paper, we propose a new unsupervised deep convolutional autoencoder-based clustering model that groups together cellular and structural morphologies of tumor in 246 ICC digitized whole slides, based on visual similarity. From this visual dictionary of histologic patterns, we use the clusters as covariates to train Cox-proportional hazard survival models. In univariate analysis, three clusters were significantly associated with recurrence-free survival. Combinations of these clusters were significant in multivariate analysis. In a multivariate analysis of all clusters, five showed significance to recurrence-free survival, however the overall model was not measured to be significant. Finally, a pathologist assigned clinical terminology to the significant clusters in the visual dictionary and found evidence supporting the hypothesis that collagen-enriched fibrosis plays a role in disease severity. These results offer insight into the future of cancer subtyping and show that computational pathology can contribute to disease prognostication, especially in rare cancers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge