Mehrtash Babadi
A Coding Theory Perspective on Multiplexed Molecular Profiling of Biological Tissues
Feb 02, 2021


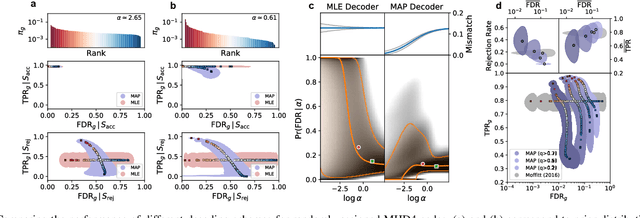
Abstract:High-throughput and quantitative experimental technologies are experiencing rapid advances in the biological sciences. One important recent technique is multiplexed fluorescence in situ hybridization (mFISH), which enables the identification and localization of large numbers of individual strands of RNA within single cells. Core to that technology is a coding problem: with each RNA sequence of interest being a codeword, how to design a codebook of probes, and how to decode the resulting noisy measurements? Published work has relied on assumptions of uniformly distributed codewords and binary symmetric channels for decoding and to a lesser degree for code construction. Here we establish that both of these assumptions are inappropriate in the context of mFISH experiments and substantial decoding performance gains can be obtained by using more appropriate, less classical, assumptions. We propose a more appropriate asymmetric channel model that can be readily parameterized from data and use it to develop a maximum a posteriori (MAP) decoders. We show that false discovery rate for rare RNAs, which is the key experimental metric, is vastly improved with MAP decoders even when employed with the existing sub-optimal codebook. Using an evolutionary optimization methodology, we further show that by permuting the codebook to better align with the prior, which is an experimentally straightforward procedure, significant further improvements are possible.
CellSegmenter: unsupervised representation learning and instance segmentation of modular images
Nov 25, 2020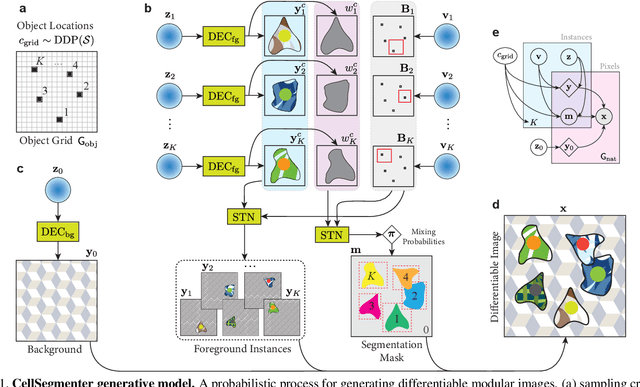
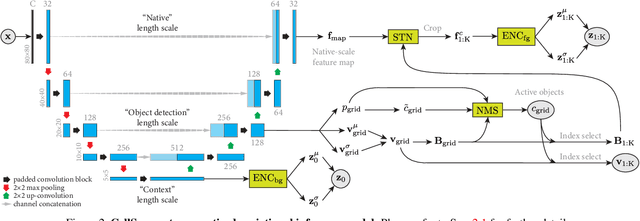
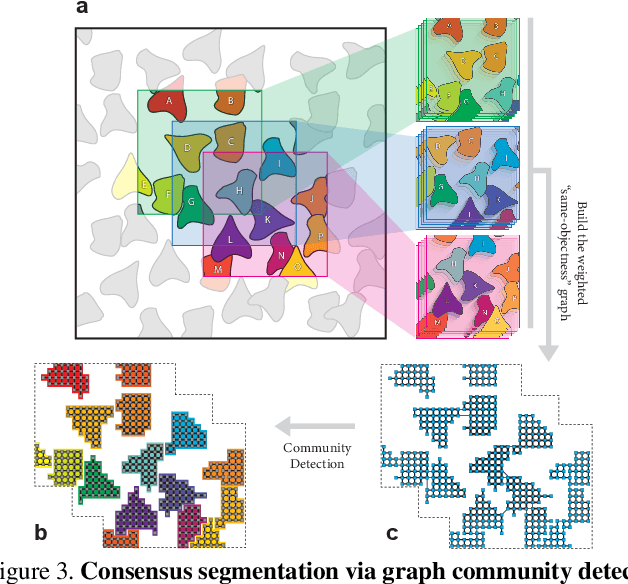
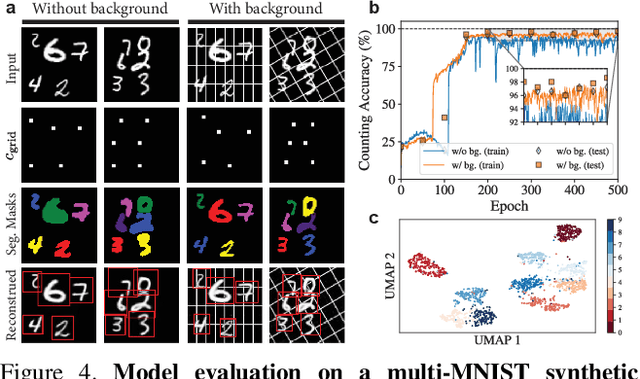
Abstract:We introduce CellSegmenter, a structured deep generative model and an amortized inference framework for unsupervised representation learning and instance segmentation tasks. The proposed inference algorithm is convolutional and parallelized, without any recurrent mechanisms, and is able to resolve object-object occlusion while simultaneously treating distant non-occluding objects independently. This leads to extremely fast training times while allowing extrapolation to arbitrary number of instances. We further introduce a transparent posterior regularization strategy that encourages scene reconstructions with fewest localized objects and a low-complexity background. We evaluate our method on a challenging synthetic multi-MNIST dataset with a structured background and achieve nearly perfect accuracy with only a few hundred training epochs. Finally, we show segmentation results obtained for a cell nuclei imaging dataset, demonstrating the ability of our method to provide high-quality segmentations while also handling realistic use cases involving large number of instances.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge