Mathieu Boudreau
Multi-Domain Data Aggregation for Axon and Myelin Segmentation in Histology Images
Sep 17, 2024
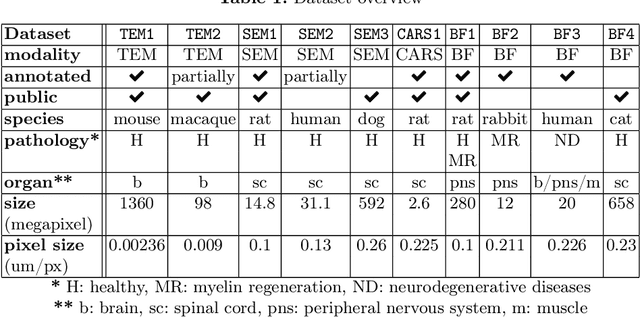
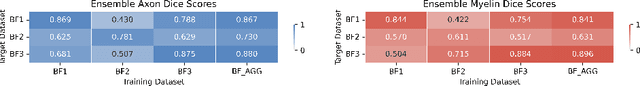
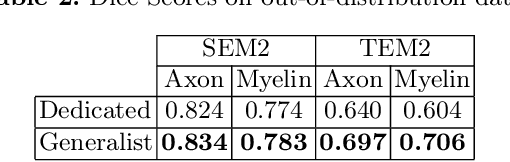
Abstract:Quantifying axon and myelin properties (e.g., axon diameter, myelin thickness, g-ratio) in histology images can provide useful information about microstructural changes caused by neurodegenerative diseases. Automatic tissue segmentation is an important tool for these datasets, as a single stained section can contain up to thousands of axons. Advances in deep learning have made this task quick and reliable with minimal overhead, but a deep learning model trained by one research group will hardly ever be usable by other groups due to differences in their histology training data. This is partly due to subject diversity (different body parts, species, genetics, pathologies) and also to the range of modern microscopy imaging techniques resulting in a wide variability of image features (i.e., contrast, resolution). There is a pressing need to make AI accessible to neuroscience researchers to facilitate and accelerate their workflow, but publicly available models are scarce and poorly maintained. Our approach is to aggregate data from multiple imaging modalities (bright field, electron microscopy, Raman spectroscopy) and species (mouse, rat, rabbit, human), to create an open-source, durable tool for axon and myelin segmentation. Our generalist model makes it easier for researchers to process their data and can be fine-tuned for better performance on specific domains. We study the benefits of different aggregation schemes. This multi-domain segmentation model performs better than single-modality dedicated learners (p=0.03077), generalizes better on out-of-distribution data and is easier to use and maintain. Importantly, we package the segmentation tool into a well-maintained open-source software ecosystem (see https://github.com/axondeepseg/axondeepseg).
Deep Active Learning for Axon-Myelin Segmentation on Histology Data
Jul 11, 2019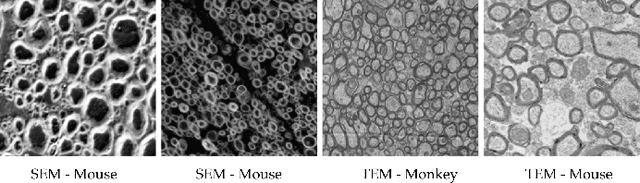
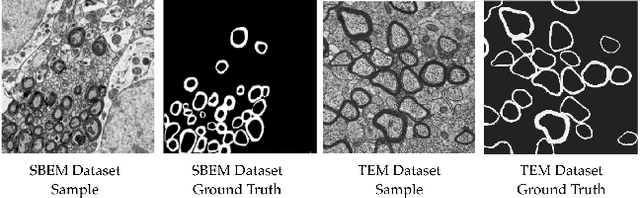
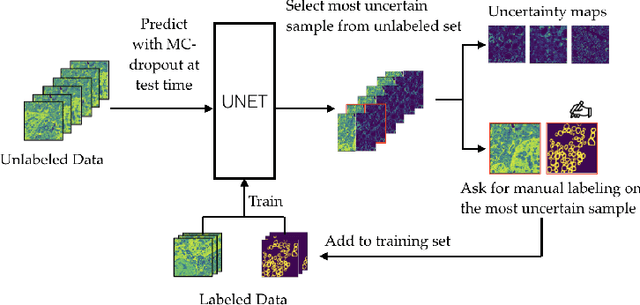
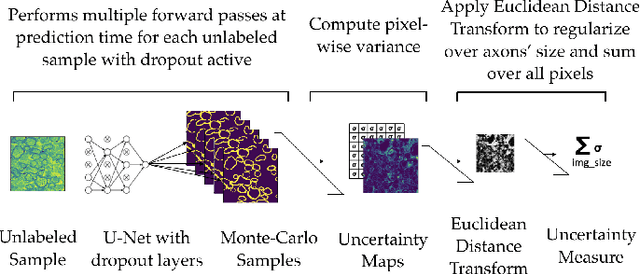

Abstract:Semantic segmentation is a crucial task in biomedical image processing, which recent breakthroughs in deep learning have allowed to improve. However, deep learning methods in general are not yet widely used in practice since they require large amount of data for training complex models. This is particularly challenging for biomedical images, because data and ground truths are a scarce resource. Annotation efforts for biomedical images come with a real cost, since experts have to manually label images at pixel-level on samples usually containing many instances of the target anatomy (e.g. in histology samples: neurons, astrocytes, mitochondria, etc.). In this paper we provide a framework for Deep Active Learning applied to a real-world scenario. Our framework relies on the U-Net architecture and overall uncertainty measure to suggest which sample to annotate. It takes advantage of the uncertainty measure obtained by taking Monte Carlo samples while using Dropout regularization scheme. Experiments were done on spinal cord and brain microscopic histology samples to perform a myelin segmentation task. Two realistic small datasets of 14 and 24 images were used, from different acquisition settings (Serial Block-Face Electron Microscopy and Transmitting Electron Microscopy) and showed that our method reached a maximum Dice value after adding 3 uncertainty-selected samples to the initial training set, versus 15 randomly-selected samples, thereby significantly reducing the annotation effort. We focused on a plausible scenario and showed evidence that this straightforward implementation achieves a high segmentation performance with very few labelled samples. We believe our framework may benefit any biomedical researcher willing to obtain fast and accurate image segmentation on their own dataset. The code is freely available at https://github.com/neuropoly/deep-active-learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge