Maryam Daniali
Language Models for Automated Classification of Brain MRI Reports and Growth Chart Generation
Mar 15, 2025Abstract:Clinically acquired brain MRIs and radiology reports are valuable but underutilized resources due to the challenges of manual analysis and data heterogeneity. We developed fine-tuned language models (LMs) to classify brain MRI reports as normal (reports with limited pathology) or abnormal, fine-tuning BERT, BioBERT, ClinicalBERT, and RadBERT on 44,661 reports. We also explored the reasoning capabilities of a leading LM, Gemini 1.5-Pro, for normal report categorization. Automated image processing and modeling generated brain growth charts from LM-classified normal scans, comparing them to human-derived charts. Fine-tuned LMs achieved high classification performance (F1-Score >97%), with unbalanced training mitigating class imbalance. Performance was robust on out-of-distribution data, with full text outperforming summary (impression) sections. Gemini 1.5-Pro showed a promising categorization performance, especially with clinical inference. LM-derived brain growth charts were nearly identical to human-annotated charts (r = 0.99, p < 2.2e-16). Our LMs offer scalable analysis of radiology reports, enabling automated classification of brain MRIs in large datasets. One application is automated generation of brain growth charts for benchmarking quantitative image features. Further research is needed to address data heterogeneity and optimize LM reasoning.
Perception Over Time: Temporal Dynamics for Robust Image Understanding
Mar 11, 2022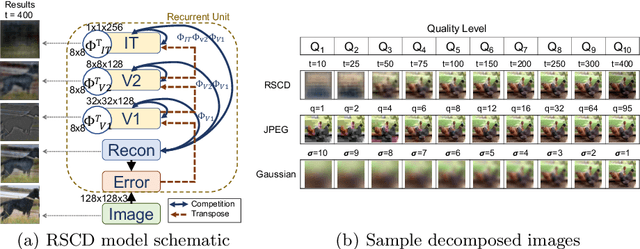
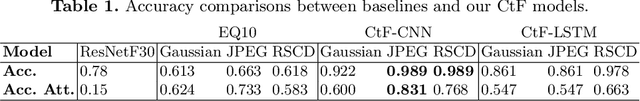
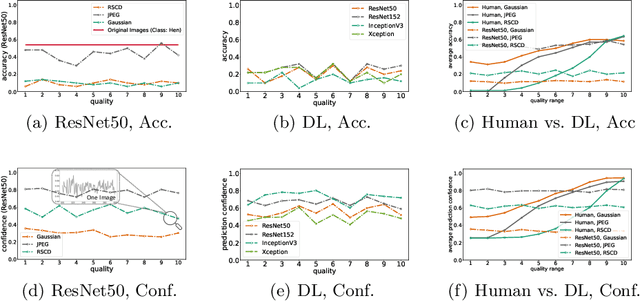
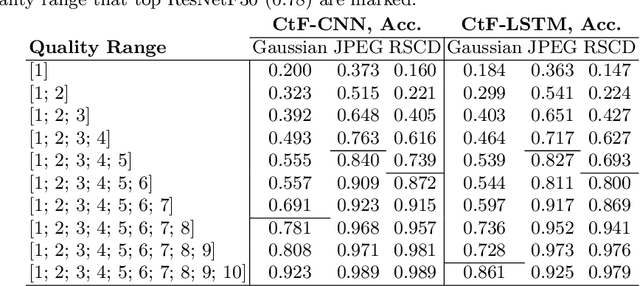
Abstract:While deep learning surpasses human-level performance in narrow and specific vision tasks, it is fragile and over-confident in classification. For example, minor transformations in perspective, illumination, or object deformation in the image space can result in drastically different labeling, which is especially transparent via adversarial perturbations. On the other hand, human visual perception is orders of magnitude more robust to changes in the input stimulus. But unfortunately, we are far from fully understanding and integrating the underlying mechanisms that result in such robust perception. In this work, we introduce a novel method of incorporating temporal dynamics into static image understanding. We describe a neuro-inspired method that decomposes a single image into a series of coarse-to-fine images that simulates how biological vision integrates information over time. Next, we demonstrate how our novel visual perception framework can utilize this information "over time" using a biologically plausible algorithm with recurrent units, and as a result, significantly improving its accuracy and robustness over standard CNNs. We also compare our proposed approach with state-of-the-art models and explicitly quantify our adversarial robustness properties through multiple ablation studies. Our quantitative and qualitative results convincingly demonstrate exciting and transformative improvements over the standard computer vision and deep learning architectures used today.
The Selectivity and Competition of the Mind's Eye in Visual Perception
Nov 23, 2020
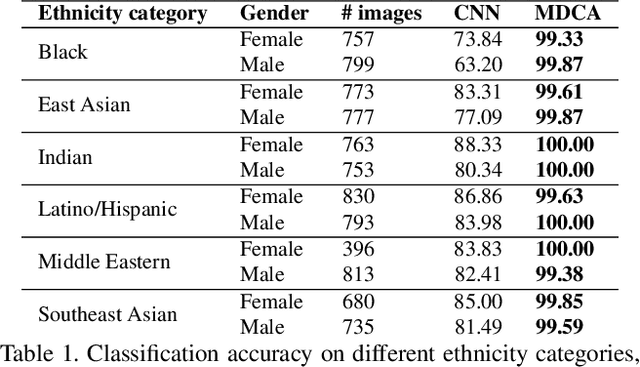
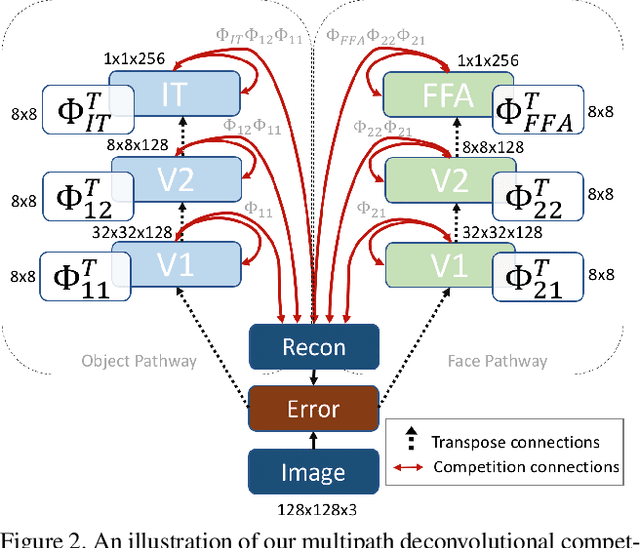

Abstract:Research has shown that neurons within the brain are selective to certain stimuli. For example, the fusiform face area (FFA) region is known by neuroscientists to selectively activate when people see faces over non-face objects. However, the mechanisms by which the primary visual system directs information to the correct higher levels of the brain are currently unknown. In our work, we advance the understanding of the neural mechanisms of perception by creating a novel computational model that incorporates lateral and top down feedback in the form of hierarchical competition. We show that these elements can help explain the information flow and selectivity of high level areas within the brain. Additionally, we present both quantitative and qualitative results that demonstrate consistency with general themes and specific responses observed in the visual system. Finally, we show that our generative framework enables a wide range of applications in computer vision, including overcoming issues of bias that have been discovered in standard deep learning models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge