Lucy Godson
National Pathology Imaging Cooperative, Leeds Teaching Hospitals NHS Trust, Leeds, UK
Machine learning-based multimodal prognostic models integrating pathology images and high-throughput omic data for overall survival prediction in cancer: a systematic review
Jul 22, 2025Abstract:Multimodal machine learning integrating histopathology and molecular data shows promise for cancer prognostication. We systematically reviewed studies combining whole slide images (WSIs) and high-throughput omics to predict overall survival. Searches of EMBASE, PubMed, and Cochrane CENTRAL (12/08/2024), plus citation screening, identified eligible studies. Data extraction used CHARMS; bias was assessed with PROBAST+AI; synthesis followed SWiM and PRISMA 2020. Protocol: PROSPERO (CRD42024594745). Forty-eight studies (all since 2017) across 19 cancer types met criteria; all used The Cancer Genome Atlas. Approaches included regularised Cox regression (n=4), classical ML (n=13), and deep learning (n=31). Reported c-indices ranged 0.550-0.857; multimodal models typically outperformed unimodal ones. However, all studies showed unclear/high bias, limited external validation, and little focus on clinical utility. Multimodal WSI-omics survival prediction is a fast-growing field with promising results but needs improved methodological rigor, broader datasets, and clinical evaluation. Funded by NPIC, Leeds Teaching Hospitals NHS Trust, UK (Project 104687), supported by UKRI Industrial Strategy Challenge Fund.
Histopathology Foundation Models Enable Accurate Ovarian Cancer Subtype Classification
May 16, 2024Abstract:Large pretrained transformers are increasingly being developed as generalised foundation models which can underpin powerful task-specific artificial intelligence models. Histopathology foundation models show promise across many tasks, but analyses have been limited by arbitrary hyperparameters that were not tuned to the specific task/dataset. We report the most rigorous single-task validation conducted to date of a histopathology foundation model, and the first performed in ovarian cancer subtyping. Attention-based multiple instance learning classifiers were compared using vision transformer and ResNet features generated through varied preprocessing and pretraining procedures. The training set consisted of 1864 whole slide images from 434 ovarian carcinoma cases at Leeds Hospitals. Five-class classification performance was evaluated through five-fold cross-validation, and these cross-validation models were ensembled for evaluation on a hold-out test set and an external set from the Transcanadian study. Reporting followed the TRIPOD+AI checklist. The vision transformer-based histopathology foundation model, UNI, performed best in every evaluation, with five-class balanced accuracies of 88% and 93% in hold-out internal and external testing, compared to the best ResNet model scores of 68% and 81%, respectively. Normalisations and augmentations aided the generalisability of ResNet-based models, but these still did not match the performance of UNI, which gave the best external performance in any ovarian cancer subtyping study to date. Histopathology foundation models offer a clear benefit to subtyping, improving classification performance to a degree where clinical utility is tangible, albeit with an increased computational burden. Such models could provide a second opinion in challenging cases and may improve the accuracy, objectivity, and efficiency of pathological diagnoses overall.
Weakly-supervised learning for image-based classification of primary melanomas into genomic immune subgroups
Feb 23, 2022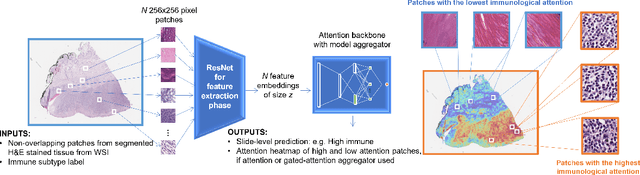

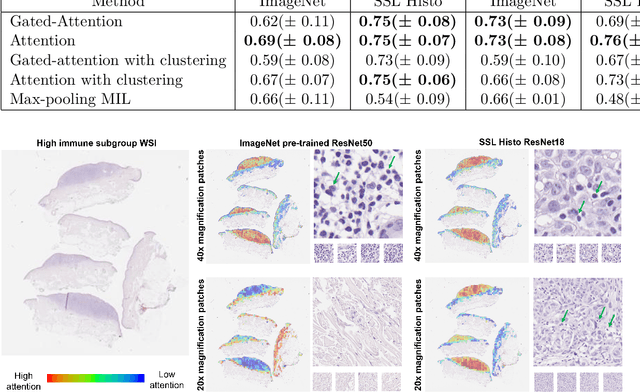
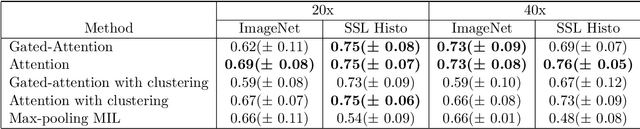
Abstract:Determining early-stage prognostic markers and stratifying patients for effective treatment are two key challenges for improving outcomes for melanoma patients. Previous studies have used tumour transcriptome data to stratify patients into immune subgroups, which were associated with differential melanoma specific survival and potential treatment strategies. However, acquiring transcriptome data is a time-consuming and costly process. Moreover, it is not routinely used in the current clinical workflow. Here we attempt to overcome this by developing deep learning models to classify gigapixel H&E stained pathology slides, which are well established in clinical workflows, into these immune subgroups. Previous subtyping approaches have employed supervised learning which requires fully annotated data, or have only examined single genetic mutations in melanoma patients. We leverage a multiple-instance learning approach, which only requires slide-level labels and uses an attention mechanism to highlight regions of high importance to the classification. Moreover, we show that pathology-specific self-supervised models generate better representations compared to pathology-agnostic models for improving our model performance, achieving a mean AUC of 0.76 for classifying histopathology images as high or low immune subgroups. We anticipate that this method may allow us to find new biomarkers of high importance and could act as a tool for clinicians to infer the immune landscape of tumours and stratify patients, without needing to carry out additional expensive genetic tests.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge