Lucian Chan
Embracing assay heterogeneity with neural processes for markedly improved bioactivity predictions
Aug 17, 2023Abstract:Predicting the bioactivity of a ligand is one of the hardest and most important challenges in computer-aided drug discovery. Despite years of data collection and curation efforts by research organizations worldwide, bioactivity data remains sparse and heterogeneous, thus hampering efforts to build predictive models that are accurate, transferable and robust. The intrinsic variability of the experimental data is further compounded by data aggregation practices that neglect heterogeneity to overcome sparsity. Here we discuss the limitations of these practices and present a hierarchical meta-learning framework that exploits the information synergy across disparate assays by successfully accounting for assay heterogeneity. We show that the model achieves a drastic improvement in affinity prediction across diverse protein targets and assay types compared to conventional baselines. It can quickly adapt to new target contexts using very few observations, thus enabling large-scale virtual screening in early-phase drug discovery.
3D pride without 2D prejudice: Bias-controlled multi-level generative models for structure-based ligand design
Apr 22, 2022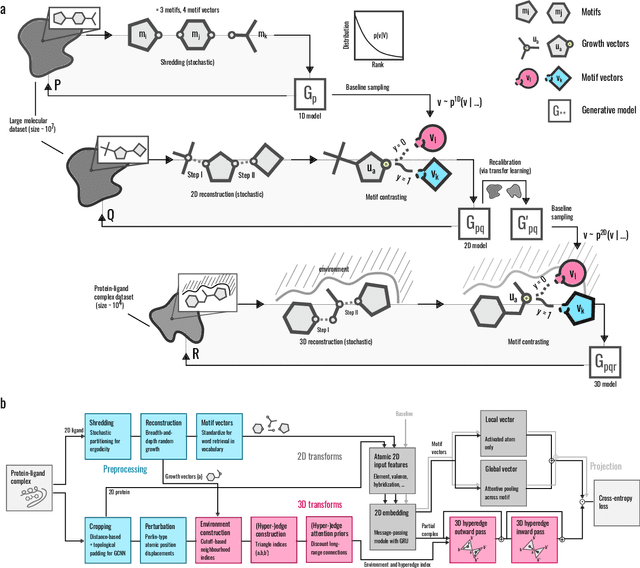
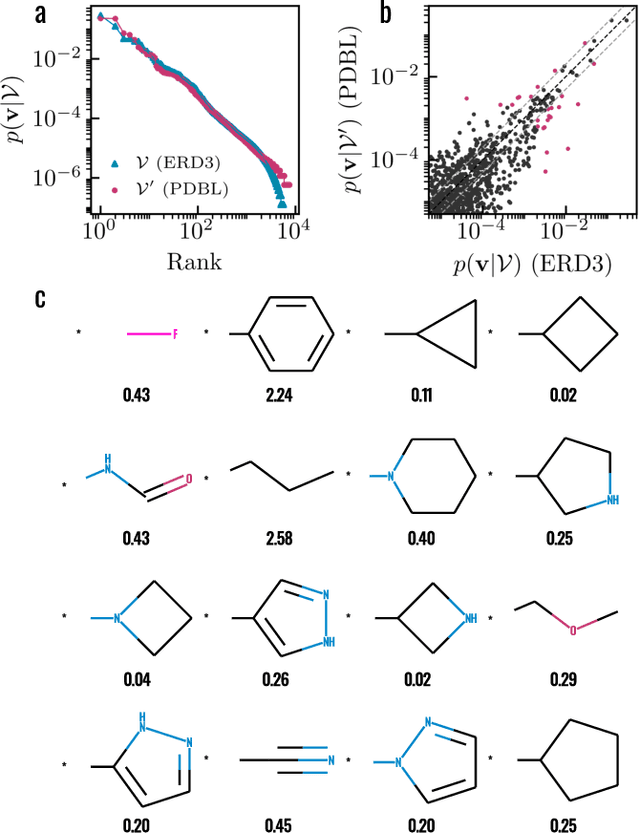
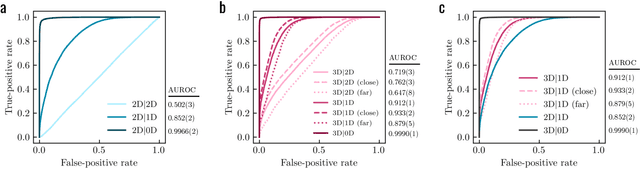
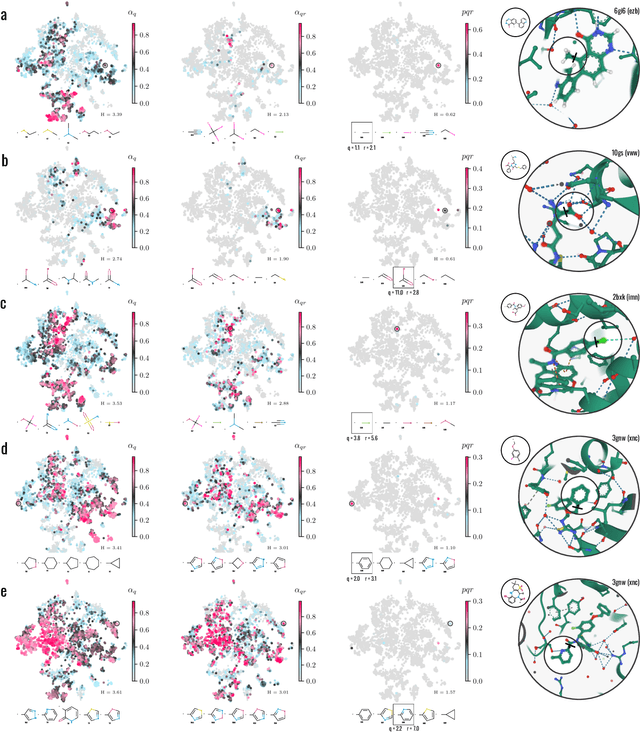
Abstract:Generative models for structure-based molecular design hold significant promise for drug discovery, with the potential to speed up the hit-to-lead development cycle, while improving the quality of drug candidates and reducing costs. Data sparsity and bias are, however, two main roadblocks to the development of 3D-aware models. Here we propose a first-in-kind training protocol based on multi-level contrastive learning for improved bias control and data efficiency. The framework leverages the large data resources available for 2D generative modelling with datasets of ligand-protein complexes. The result are hierarchical generative models that are topologically unbiased, explainable and customizable. We show how, by deconvolving the generative posterior into chemical, topological and structural context factors, we not only avoid common pitfalls in the design and evaluation of generative models, but furthermore gain detailed insight into the generative process itself. This improved transparency significantly aids method development, besides allowing fine-grained control over novelty vs familiarity.
Noise Contrastive Meta-Learning for Conditional Density Estimation using Kernel Mean Embeddings
Jun 05, 2019
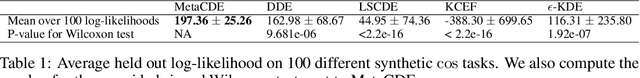
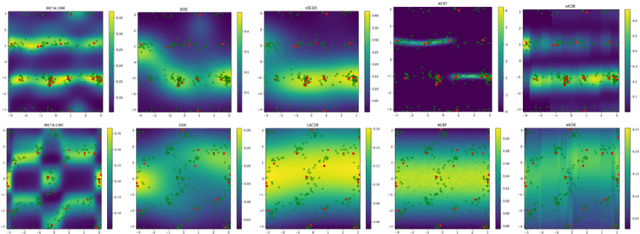

Abstract:Current meta-learning approaches focus on learning functional representations of relationships between variables, i.e. on estimating conditional expectations in regression. In many applications, however, we are faced with conditional distributions which cannot be meaningfully summarized using expectation only (due to e.g. multimodality). Hence, we consider the problem of conditional density estimation in the meta-learning setting. We introduce a novel technique for meta-learning which combines neural representation and noise-contrastive estimation with the established literature of conditional mean embeddings into reproducing kernel Hilbert spaces. The method is validated on synthetic and real-world problems, demonstrating the utility of sharing learned representations across multiple conditional density estimation tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge