Lingsheng Cai
Deep Molecular Representation Learning via Fusing Physical and Chemical Information
Nov 28, 2021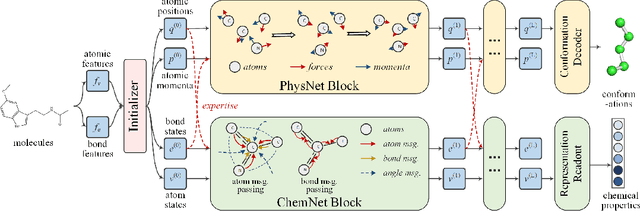
Abstract:Molecular representation learning is the first yet vital step in combining deep learning and molecular science. To push the boundaries of molecular representation learning, we present PhysChem, a novel neural architecture that learns molecular representations via fusing physical and chemical information of molecules. PhysChem is composed of a physicist network (PhysNet) and a chemist network (ChemNet). PhysNet is a neural physical engine that learns molecular conformations through simulating molecular dynamics with parameterized forces; ChemNet implements geometry-aware deep message-passing to learn chemical / biomedical properties of molecules. Two networks specialize in their own tasks and cooperate by providing expertise to each other. By fusing physical and chemical information, PhysChem achieved state-of-the-art performances on MoleculeNet, a standard molecular machine learning benchmark. The effectiveness of PhysChem was further corroborated on cutting-edge datasets of SARS-CoV-2.
HamNet: Conformation-Guided Molecular Representation with Hamiltonian Neural Networks
May 08, 2021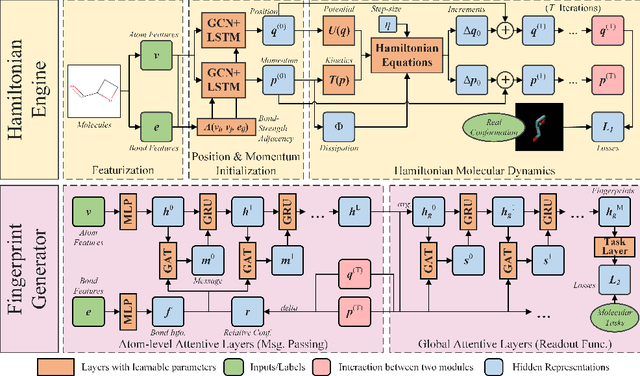
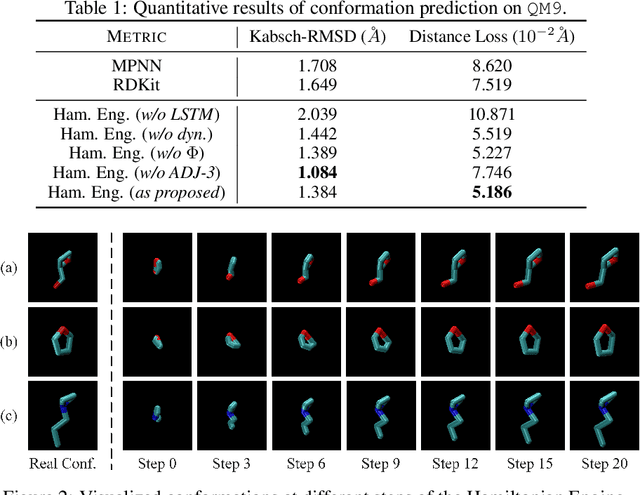
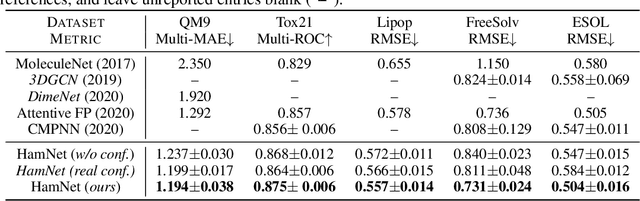
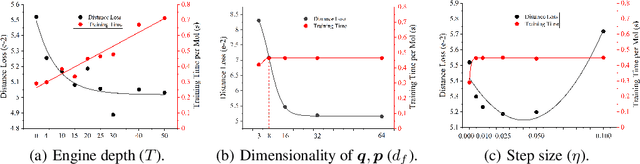
Abstract:Well-designed molecular representations (fingerprints) are vital to combine medical chemistry and deep learning. Whereas incorporating 3D geometry of molecules (i.e. conformations) in their representations seems beneficial, current 3D algorithms are still in infancy. In this paper, we propose a novel molecular representation algorithm which preserves 3D conformations of molecules with a Molecular Hamiltonian Network (HamNet). In HamNet, implicit positions and momentums of atoms in a molecule interact in the Hamiltonian Engine following the discretized Hamiltonian equations. These implicit coordinations are supervised with real conformations with translation- & rotation-invariant losses, and further used as inputs to the Fingerprint Generator, a message-passing neural network. Experiments show that the Hamiltonian Engine can well preserve molecular conformations, and that the fingerprints generated by HamNet achieve state-of-the-art performances on MoleculeNet, a standard molecular machine learning benchmark.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge