Leila Saadatifard
DVNet: A Memory-Efficient Three-Dimensional CNN for Large-Scale Neurovascular Reconstruction
Feb 04, 2020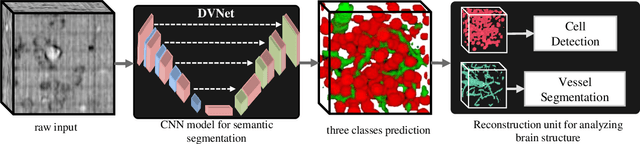
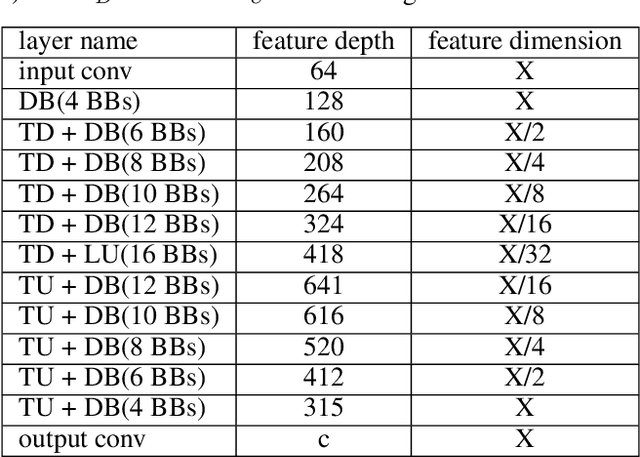
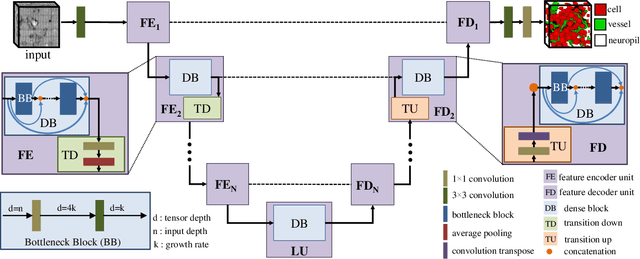

Abstract:Maps of brain microarchitecture are important for understanding neurological function and behavior, including alterations caused by chronic conditions such as neurodegenerative disease. Techniques such as knife-edge scanning microscopy (KESM) provide the potential for whole organ imaging at sub-cellular resolution. However, multi-terabyte data sizes make manual annotation impractical and automatic segmentation challenging. Densely packed cells combined with interconnected microvascular networks are a challenge for current segmentation algorithms. The massive size of high-throughput microscopy data necessitates fast and largely unsupervised algorithms. In this paper, we investigate a fully-convolutional, deep, and densely-connected encoder-decoder for pixel-wise semantic segmentation. The excessive memory complexity often encountered with deep and dense networks is mitigated using skip connections, resulting in fewer parameters and enabling a significant performance increase over prior architectures. The proposed network provides superior performance for semantic segmentation problems applied to open-source benchmarks. We finally demonstrate our network for cellular and microvascular segmentation, enabling quantitative metrics for organ-scale neurovascular analysis.
Three-Dimensional GPU-Accelerated Active Contours for Automated Localization of Cells in Large Images
Apr 17, 2018



Abstract:Cell segmentation in microscopy is a challenging problem, since cells are often asymmetric and densely packed. This becomes particularly challenging for extremely large images, since manual intervention and processing time can make segmentation intractable. In this paper, we present an efficient and highly parallel formulation for symmetric three-dimensional (3D) contour evolution that extends previous work on fast two-dimensional active contours. We provide a formulation for optimization on 3D images, as well as a strategy for accelerating computation on consumer graphics hardware. The proposed software takes advantage of Monte-Carlo sampling schemes in order to speed up convergence and reduce thread divergence. Experimental results show that this method provides superior performance for large 2D and 3D cell segmentation tasks when compared to existing methods on large 3D brain images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge