Khader Shameer
TrialGraph: Machine Intelligence Enabled Insight from Graph Modelling of Clinical Trials
Dec 15, 2021
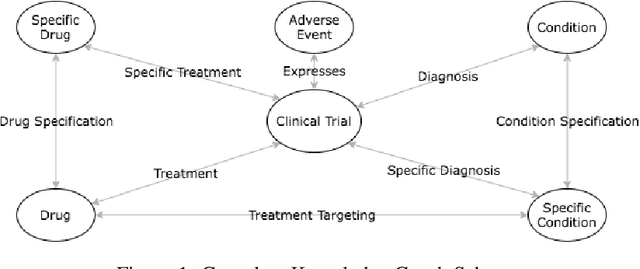
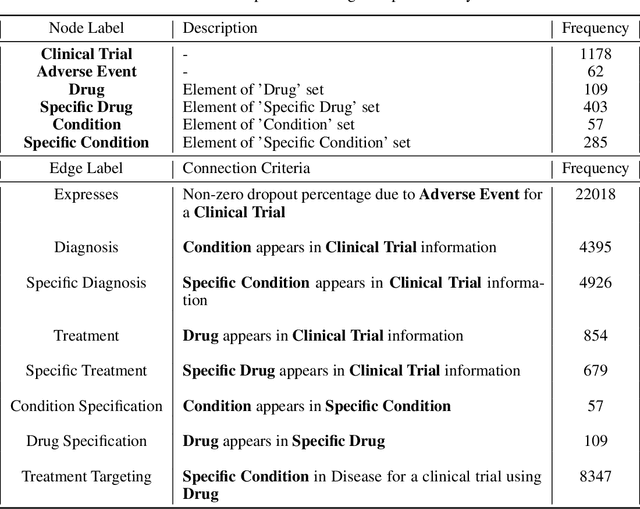

Abstract:A major impediment to successful drug development is the complexity, cost, and scale of clinical trials. The detailed internal structure of clinical trial data can make conventional optimization difficult to achieve. Recent advances in machine learning, specifically graph-structured data analysis, have the potential to enable significant progress in improving the clinical trial design. TrialGraph seeks to apply these methodologies to produce a proof-of-concept framework for developing models which can aid drug development and benefit patients. In this work, we first introduce a curated clinical trial data set compiled from the CT.gov, AACT and TrialTrove databases (n=1191 trials; representing one million patients) and describe the conversion of this data to graph-structured formats. We then detail the mathematical basis and implementation of a selection of graph machine learning algorithms, which typically use standard machine classifiers on graph data embedded in a low-dimensional feature space. We trained these models to predict side effect information for a clinical trial given information on the disease, existing medical conditions, and treatment. The MetaPath2Vec algorithm performed exceptionally well, with standard Logistic Regression, Decision Tree, Random Forest, Support Vector, and Neural Network classifiers exhibiting typical ROC-AUC scores of 0.85, 0.68, 0.86, 0.80, and 0.77, respectively. Remarkably, the best performing classifiers could only produce typical ROC-AUC scores of 0.70 when trained on equivalent array-structured data. Our work demonstrates that graph modelling can significantly improve prediction accuracy on appropriate datasets. Successive versions of the project that refine modelling assumptions and incorporate more data types can produce excellent predictors with real-world applications in drug development.
Mammography Assessment using Multi-Scale Deep Classifiers
Jun 30, 2018
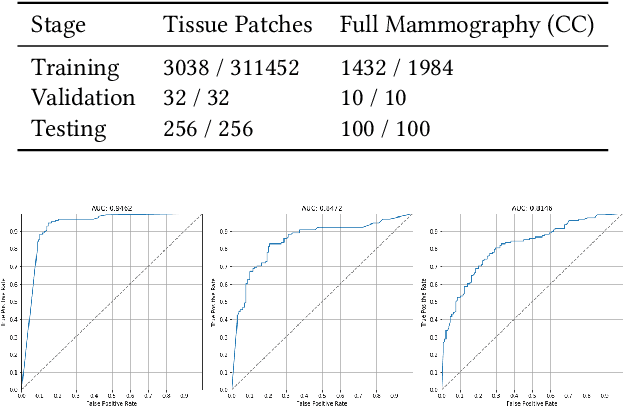
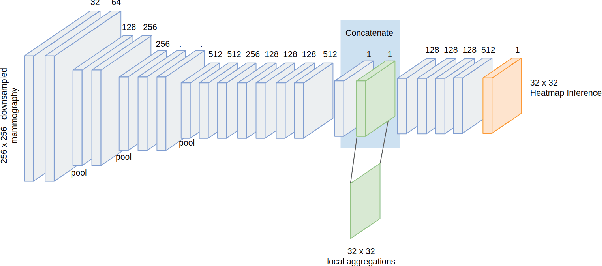
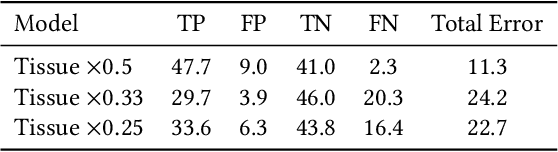
Abstract:Applying deep learning methods to mammography assessment has remained a challenging topic. Dense noise with sparse expressions, mega-pixel raw data resolution, lack of diverse examples have all been factors affecting performance. The lack of pixel-level ground truths have especially limited segmentation methods in pushing beyond approximately bounding regions. We propose a classification approach grounded in high performance tissue assessment as an alternative to all-in-one localization and assessment models that is also capable of pinpointing the causal pixels. First, the objective of the mammography assessment task is formalized in the context of local tissue classifiers. Then, the accuracy of a convolutional neural net is evaluated on classifying patches of tissue with suspicious findings at varying scales, where highest obtained AUC is above $0.9$. The local evaluations of one such expert tissue classifier is used to augment the results of a heatmap regression model and additionally recover the exact causal regions at high resolution as a saliency image suitable for clinical settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge