José Luis Rojo-Álvarez
Latent Diffusion for Internet of Things Attack Data Generation in Intrusion Detection
Jan 23, 2026Abstract:Intrusion Detection Systems (IDSs) are a key component for protecting Internet of Things (IoT) environments. However, in Machine Learning-based (ML-based) IDSs, performance is often degraded by the strong class imbalance between benign and attack traffic. Although data augmentation has been widely explored to mitigate this issue, existing approaches typically rely on simple oversampling techniques or generative models that struggle to simultaneously achieve high sample fidelity, diversity, and computational efficiency. To address these limitations, we propose the use of a Latent Diffusion Model (LDM) for attack data augmentation in IoT intrusion detection and provide a comprehensive comparison against state-of-the-art baselines. Experiments were conducted on three representative IoT attack types, specifically Distributed Denial-of-Service (DDoS), Mirai, and Man-in-the-Middle, evaluating both downstream IDS performance and intrinsic generative quality using distributional, dependency-based, and diversity metrics. Results show that balancing the training data with LDM-generated samples substantially improves IDS performance, achieving F1-scores of up to 0.99 for DDoS and Mirai attacks and consistently outperforming competing methods. Additionally, quantitative and qualitative analyses demonstrate that LDMs effectively preserve feature dependencies while generating diverse samples and reduce sampling time by approximately 25\% compared to diffusion models operating directly in data space. These findings highlight latent diffusion as an effective and scalable solution for synthetic IoT attack data generation, substantially mitigating the impact of class imbalance in ML-based IDSs for IoT scenarios.
Explainable Autoencoder-Based Anomaly Detection in IEC 61850 GOOSE Networks
Jan 14, 2026Abstract:The IEC 61850 Generic Object-Oriented Substation Event (GOOSE) protocol plays a critical role in real-time protection and automation of digital substations, yet its lack of native security mechanisms can expose power systems to sophisticated cyberattacks. Traditional rule-based and supervised intrusion detection techniques struggle to detect protocol-compliant and zero-day attacks under significant class imbalance and limited availability of labeled data. This paper proposes an explainable, unsupervised multi-view anomaly detection framework for IEC 61850 GOOSE networks that explicitly separates semantic integrity and temporal availability. The approach employs asymmetric autoencoders trained only on real operational GOOSE traffic to learn distinct latent representations of sequence-based protocol semantics and timing-related transmission dynamics in normal traffic. Anomaly detection is implemented using reconstruction errors mixed with statistically grounded thresholds, enabling robust detection without specified attack types. Feature-level reconstruction analysis provides intrinsic explainability by directly linking detection outcomes to IEC 61850 protocol characteristics. The proposed framework is evaluated using real substation traffic for training and a public dataset containing normal traffic and message suppression, data manipulation, and denial-of-service attacks for testing. Experimental results show attack detection rates above 99% with false positives remaining below 5% of total traffic, demonstrating strong generalization across environments and effective operation under extreme class imbalance and interpretable anomaly attribution.
Multivariate Feature Selection and Autoencoder Embeddings of Ovarian Cancer Clinical and Genetic Data
Jan 27, 2025Abstract:This study explores a data-driven approach to discovering novel clinical and genetic markers in ovarian cancer (OC). Two main analyses were performed: (1) a nonlinear examination of an OC dataset using autoencoders, which compress data into a 3-dimensional latent space to detect potential intrinsic separability between platinum-sensitive and platinum-resistant groups; and (2) an adaptation of the informative variable identifier (IVI) to determine which features (clinical or genetic) are most relevant to disease progression. In the autoencoder analysis, a clearer pattern emerged when using clinical features and the combination of clinical and genetic data, indicating that disease progression groups can be distinguished more effectively after supervised fine tuning. For genetic data alone, this separability was less apparent but became more pronounced with a supervised approach. Using the IVI-based feature selection, key clinical variables (such as type of surgery and neoadjuvant chemotherapy) and certain gene mutations showed strong relevance, along with low-risk genetic factors. These findings highlight the strength of combining machine learning tools (autoencoders) with feature selection methods (IVI) to gain insights into ovarian cancer progression. They also underscore the potential for identifying new biomarkers that integrate clinical and genomic indicators, ultimately contributing to improved patient stratification and personalized treatment strategies.
A Unified SVM Framework for Signal Estimation
Nov 21, 2013
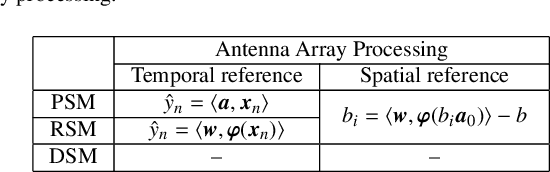
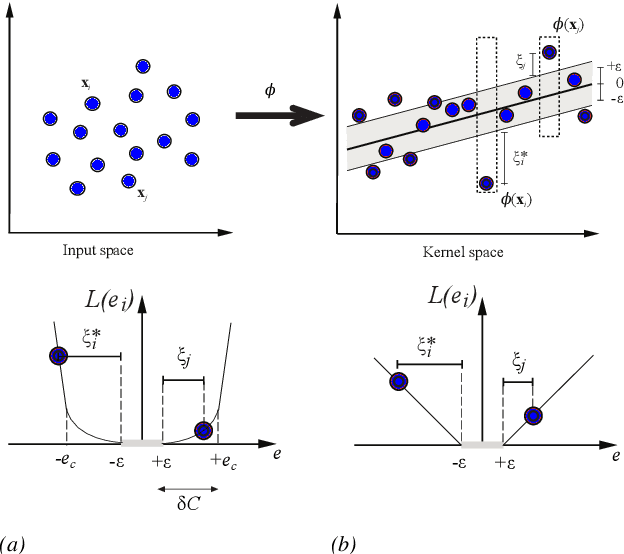

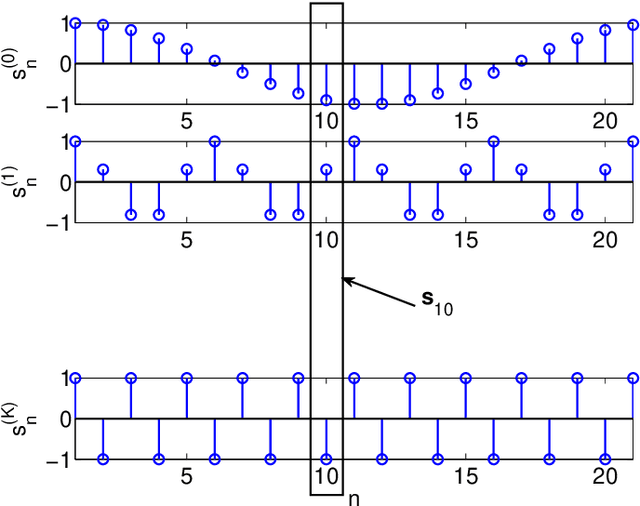
Abstract:This paper presents a unified framework to tackle estimation problems in Digital Signal Processing (DSP) using Support Vector Machines (SVMs). The use of SVMs in estimation problems has been traditionally limited to its mere use as a black-box model. Noting such limitations in the literature, we take advantage of several properties of Mercer's kernels and functional analysis to develop a family of SVM methods for estimation in DSP. Three types of signal model equations are analyzed. First, when a specific time-signal structure is assumed to model the underlying system that generated the data, the linear signal model (so called Primal Signal Model formulation) is first stated and analyzed. Then, non-linear versions of the signal structure can be readily developed by following two different approaches. On the one hand, the signal model equation is written in reproducing kernel Hilbert spaces (RKHS) using the well-known RKHS Signal Model formulation, and Mercer's kernels are readily used in SVM non-linear algorithms. On the other hand, in the alternative and not so common Dual Signal Model formulation, a signal expansion is made by using an auxiliary signal model equation given by a non-linear regression of each time instant in the observed time series. These building blocks can be used to generate different novel SVM-based methods for problems of signal estimation, and we deal with several of the most important ones in DSP. We illustrate the usefulness of this methodology by defining SVM algorithms for linear and non-linear system identification, spectral analysis, nonuniform interpolation, sparse deconvolution, and array processing. The performance of the developed SVM methods is compared to standard approaches in all these settings. The experimental results illustrate the generality, simplicity, and capabilities of the proposed SVM framework for DSP.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge