Joeran Bosma
EvalBlocks: A Modular Pipeline for Rapidly Evaluating Foundation Models in Medical Imaging
Jan 07, 2026Abstract:Developing foundation models in medical imaging requires continuous monitoring of downstream performance. Researchers are burdened with tracking numerous experiments, design choices, and their effects on performance, often relying on ad-hoc, manual workflows that are inherently slow and error-prone. We introduce EvalBlocks, a modular, plug-and-play framework for efficient evaluation of foundation models during development. Built on Snakemake, EvalBlocks supports seamless integration of new datasets, foundation models, aggregation methods, and evaluation strategies. All experiments and results are tracked centrally and are reproducible with a single command, while efficient caching and parallel execution enable scalable use on shared compute infrastructure. Demonstrated on five state-of-the-art foundation models and three medical imaging classification tasks, EvalBlocks streamlines model evaluation, enabling researchers to iterate faster and focus on model innovation rather than evaluation logistics. The framework is released as open source software at https://github.com/DIAGNijmegen/eval-blocks.
Uncertainty-Aware Semi-Supervised Learning for Prostate MRI Zonal Segmentation
May 10, 2023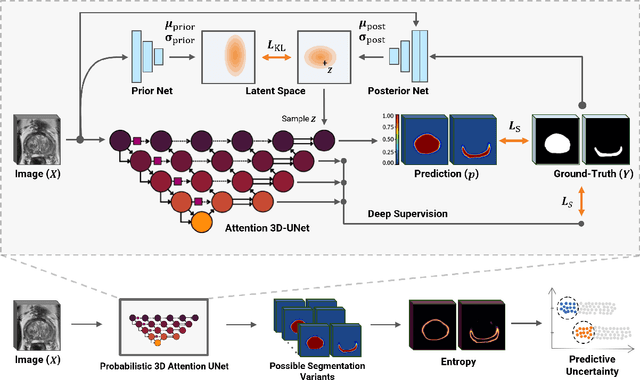
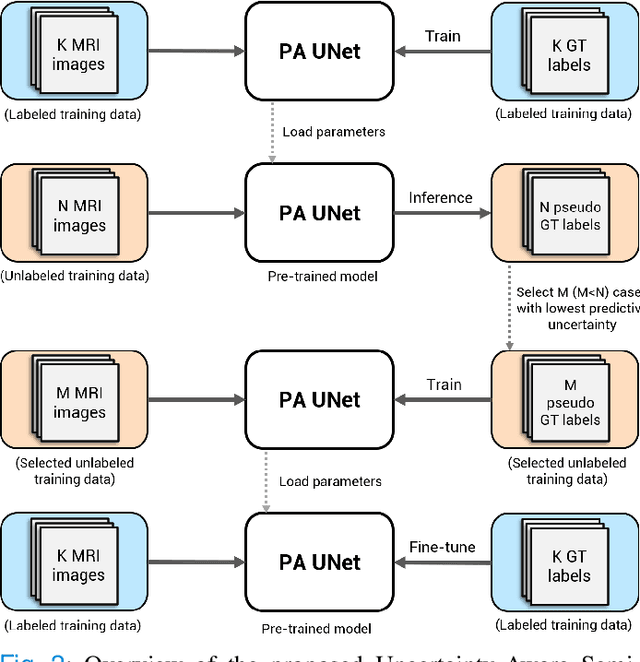
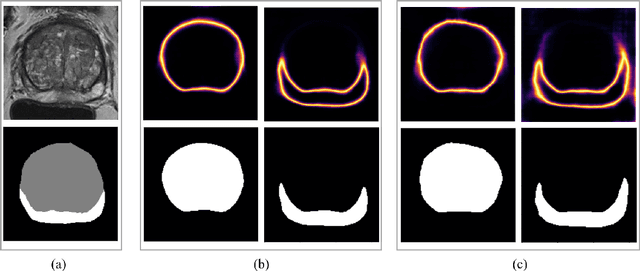
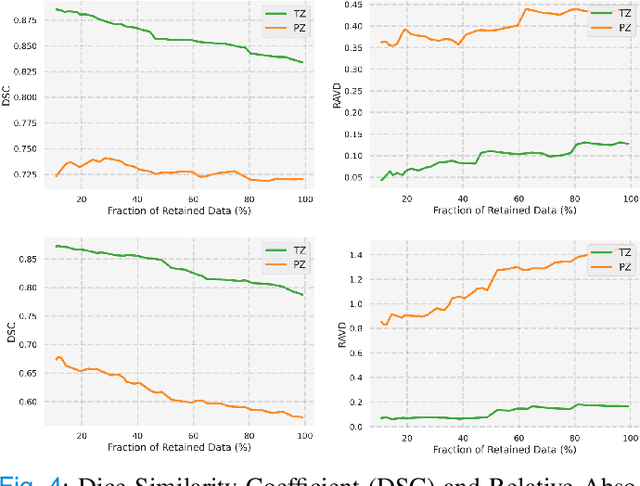
Abstract:Quality of deep convolutional neural network predictions strongly depends on the size of the training dataset and the quality of the annotations. Creating annotations, especially for 3D medical image segmentation, is time-consuming and requires expert knowledge. We propose a novel semi-supervised learning (SSL) approach that requires only a relatively small number of annotations while being able to use the remaining unlabeled data to improve model performance. Our method uses a pseudo-labeling technique that employs recent deep learning uncertainty estimation models. By using the estimated uncertainty, we were able to rank pseudo-labels and automatically select the best pseudo-annotations generated by the supervised model. We applied this to prostate zonal segmentation in T2-weighted MRI scans. Our proposed model outperformed the semi-supervised model in experiments with the ProstateX dataset and an external test set, by leveraging only a subset of unlabeled data rather than the full collection of 4953 cases, our proposed model demonstrated improved performance. The segmentation dice similarity coefficient in the transition zone and peripheral zone increased from 0.835 and 0.727 to 0.852 and 0.751, respectively, for fully supervised model and the uncertainty-aware semi-supervised learning model (USSL). Our USSL model demonstrates the potential to allow deep learning models to be trained on large datasets without requiring full annotation. Our code is available at https://github.com/DIAGNijmegen/prostateMR-USSL.
Anatomical and Diagnostic Bayesian Segmentation in Prostate MRI $-$Should Different Clinical Objectives Mandate Different Loss Functions?
Oct 25, 2021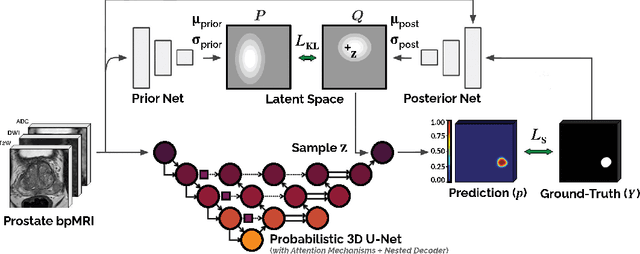
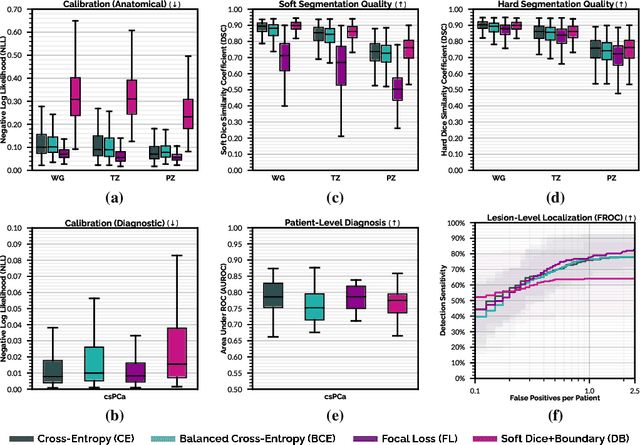
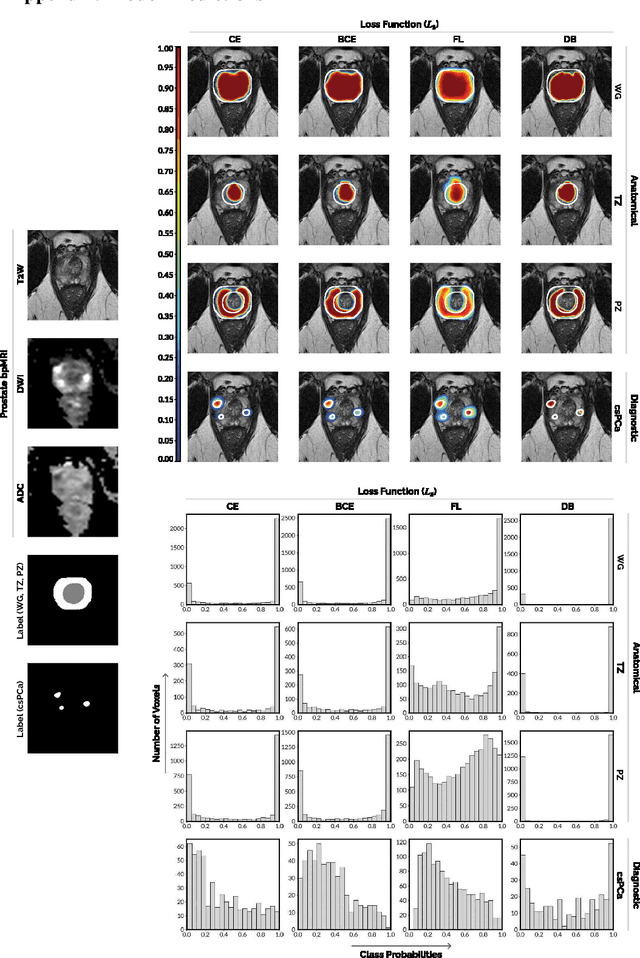
Abstract:We hypothesize that probabilistic voxel-level classification of anatomy and malignancy in prostate MRI, although typically posed as near-identical segmentation tasks via U-Nets, require different loss functions for optimal performance due to inherent differences in their clinical objectives. We investigate distribution, region and boundary-based loss functions for both tasks across 200 patient exams from the publicly-available ProstateX dataset. For evaluation, we conduct a thorough comparative analysis of model predictions and calibration, measured with respect to multi-class volume segmentation of the prostate anatomy (whole-gland, transitional zone, peripheral zone), as well as, patient-level diagnosis and lesion-level detection of clinically significant prostate cancer. Notably, we find that distribution-based loss functions (in particular, focal loss) are well-suited for diagnostic or panoptic segmentation tasks such as lesion detection, primarily due to their implicit property of inducing better calibration. Meanwhile, (with the exception of focal loss) both distribution and region/boundary-based loss functions perform equally well for anatomical or semantic segmentation tasks, such as quantification of organ shape, size and boundaries.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge