Jianan Nie
FlowMS: Flow Matching for De Novo Structure Elucidation from Mass Spectra
Mar 19, 2026Abstract:Mass spectrometry (MS) stands as a cornerstone analytical technique for molecular identification, yet de novo structure elucidation from spectra remains challenging due to the combinatorial complexity of chemical space and the inherent ambiguity of spectral fragmentation patterns. Recent deep learning approaches, including autoregressive sequence models, scaffold-based methods, and graph diffusion models, have made progress. However, diffusion-based generation for this task remains computationally demanding. Meanwhile, discrete flow matching, which has shown strong performance for graph generation, has not yet been explored for spectrum-conditioned structure elucidation. In this work, we introduce FlowMS, the first discrete flow matching framework for spectrum-conditioned de novo molecular generation. FlowMS generates molecular graphs through iterative refinement in probability space, enforcing chemical formula constraints while conditioning on spectral embeddings from a pretrained formula transformer encoder. Notably, it achieves state-of-the-art performance on 5 out of 6 metrics on the NPLIB1 benchmark: 9.15% top-1 accuracy (9.7% relative improvement over DiffMS) and 7.96 top-10 MCES (4.2% improvement over MS-BART). We also visualize the generated molecules, which further demonstrate that FlowMS produces structurally plausible candidates closely resembling ground truth structures. These results establish discrete flow matching as a promising paradigm for mass spectrometry-based structure elucidation in metabolomics and natural product discovery.
ReGNet: Reciprocal Space-Aware Long-Range Modeling and Multi-Property Prediction for Crystals
Feb 04, 2025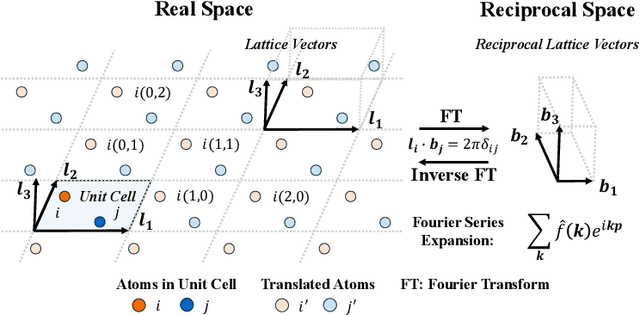
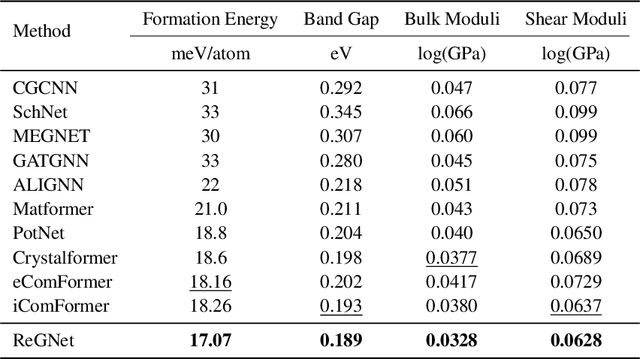
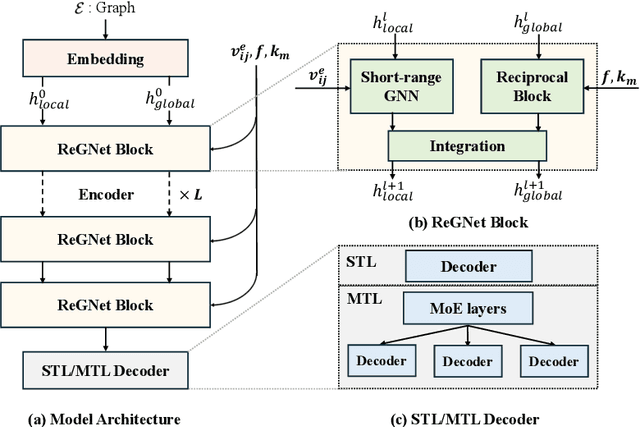
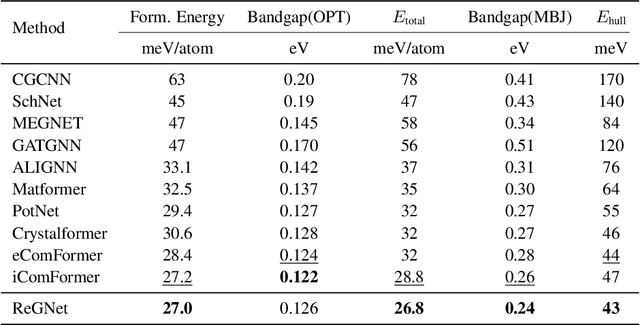
Abstract:Predicting properties of crystals from their structures is a fundamental yet challenging task in materials science. Unlike molecules, crystal structures exhibit infinite periodic arrangements of atoms, requiring methods capable of capturing both local and global information effectively. However, most current works fall short of capturing long-range interactions within periodic structures. To address this limitation, we leverage reciprocal space to efficiently encode long-range interactions with learnable filters within Fourier transforms. We introduce Reciprocal Geometry Network (ReGNet), a novel architecture that integrates geometric GNNs and reciprocal blocks to model short-range and long-range interactions, respectively. Additionally, we introduce ReGNet-MT, a multi-task extension that employs mixture of experts (MoE) for multi-property prediction. Experimental results on the JARVIS and Materials Project benchmarks demonstrate that ReGNet achieves significant performance improvements. Moreover, ReGNet-MT attains state-of-the-art results on two bandgap properties due to positive transfer, while maintaining high computational efficiency. These findings highlight the potential of our model as a scalable and accurate solution for crystal property prediction. The code will be released upon paper acceptance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge