Javier Béjar
Data Augmentation with Diffusion Models for Colon Polyp Localization on the Low Data Regime: How much real data is enough?
Nov 28, 2024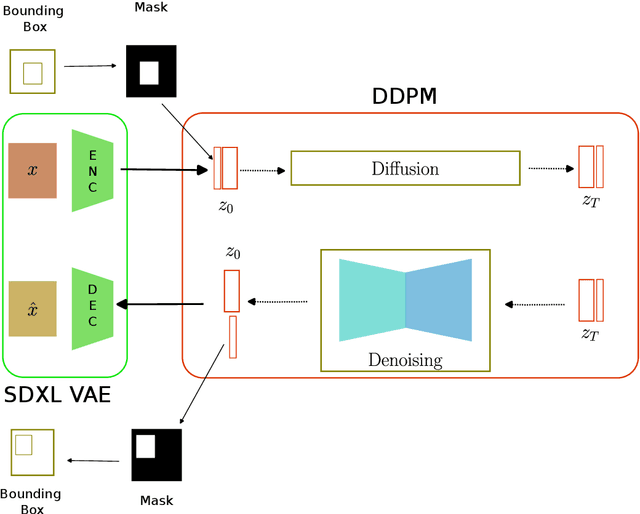
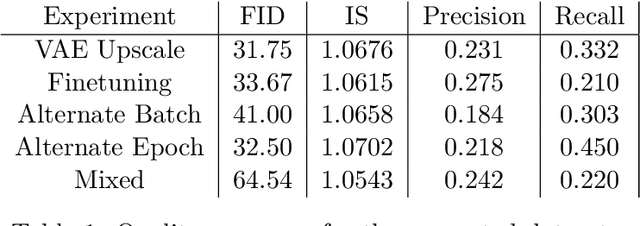
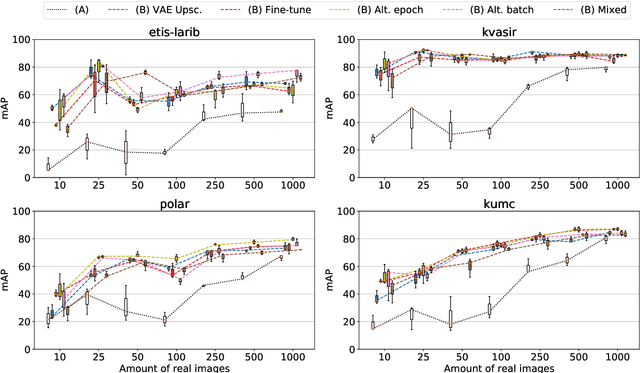
Abstract:The scarcity of data in medical domains hinders the performance of Deep Learning models. Data augmentation techniques can alleviate that problem, but they usually rely on functional transformations of the data that do not guarantee to preserve the original tasks. To approximate the distribution of the data using generative models is a way of reducing that problem and also to obtain new samples that resemble the original data. Denoising Diffusion models is a promising Deep Learning technique that can learn good approximations of different kinds of data like images, time series or tabular data. Automatic colonoscopy analysis and specifically Polyp localization in colonoscopy videos is a task that can assist clinical diagnosis and treatment. The annotation of video frames for training a deep learning model is a time consuming task and usually only small datasets can be obtained. The fine tuning of application models using a large dataset of generated data could be an alternative to improve their performance. We conduct a set of experiments training different diffusion models that can generate jointly colonoscopy images with localization annotations using a combination of existing open datasets. The generated data is used on various transfer learning experiments in the task of polyp localization with a model based on YOLO v9 on the low data regime.
Synthetic ECG Generation for Data Augmentation and Transfer Learning in Arrhythmia Classification
Nov 27, 2024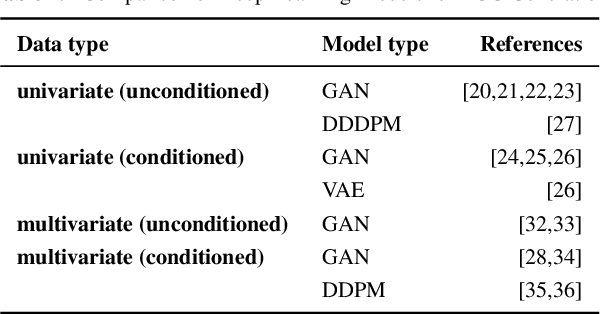
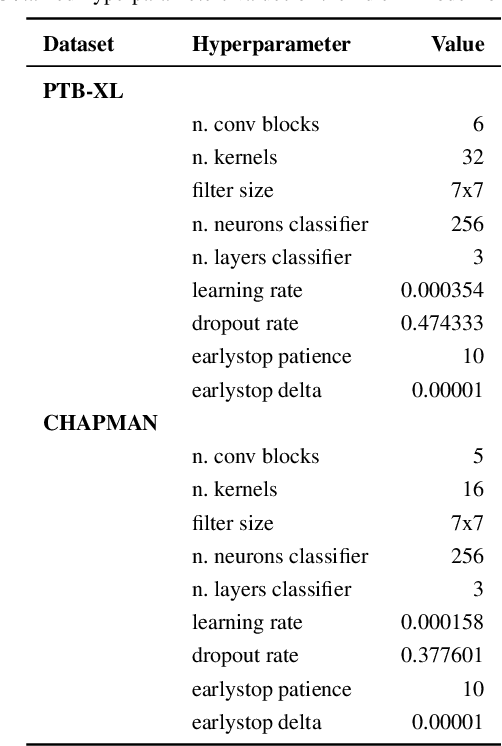



Abstract:Deep learning models need a sufficient amount of data in order to be able to find the hidden patterns in it. It is the purpose of generative modeling to learn the data distribution, thus allowing us to sample more data and augment the original dataset. In the context of physiological data, and more specifically electrocardiogram (ECG) data, given its sensitive nature and expensive data collection, we can exploit the benefits of generative models in order to enlarge existing datasets and improve downstream tasks, in our case, classification of heart rhythm. In this work, we explore the usefulness of synthetic data generated with different generative models from Deep Learning namely Diffweave, Time-Diffusion and Time-VQVAE in order to obtain better classification results for two open source multivariate ECG datasets. Moreover, we also investigate the effects of transfer learning, by fine-tuning a synthetically pre-trained model and then progressively adding increasing proportions of real data. We conclude that although the synthetic samples resemble the real ones, the classification improvement when simply augmenting the real dataset is barely noticeable on individual datasets, but when both datasets are merged the results show an increase across all metrics for the classifiers when using synthetic samples as augmented data. From the fine-tuning results the Time-VQVAE generative model has shown to be superior to the others but not powerful enough to achieve results close to a classifier trained with real data only. In addition, methods and metrics for measuring closeness between synthetic data and the real one have been explored as a side effect of the main research questions of this study.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge