Heather C. Whalley
DenseNet and Support Vector Machine classifications of major depressive disorder using vertex-wise cortical features
Nov 18, 2023



Abstract:Major depressive disorder (MDD) is a complex psychiatric disorder that affects the lives of hundreds of millions of individuals around the globe. Even today, researchers debate if morphological alterations in the brain are linked to MDD, likely due to the heterogeneity of this disorder. The application of deep learning tools to neuroimaging data, capable of capturing complex non-linear patterns, has the potential to provide diagnostic and predictive biomarkers for MDD. However, previous attempts to demarcate MDD patients and healthy controls (HC) based on segmented cortical features via linear machine learning approaches have reported low accuracies. In this study, we used globally representative data from the ENIGMA-MDD working group containing an extensive sample of people with MDD (N=2,772) and HC (N=4,240), which allows a comprehensive analysis with generalizable results. Based on the hypothesis that integration of vertex-wise cortical features can improve classification performance, we evaluated the classification of a DenseNet and a Support Vector Machine (SVM), with the expectation that the former would outperform the latter. As we analyzed a multi-site sample, we additionally applied the ComBat harmonization tool to remove potential nuisance effects of site. We found that both classifiers exhibited close to chance performance (balanced accuracy DenseNet: 51%; SVM: 53%), when estimated on unseen sites. Slightly higher classification performance (balanced accuracy DenseNet: 58%; SVM: 55%) was found when the cross-validation folds contained subjects from all sites, indicating site effect. In conclusion, the integration of vertex-wise morphometric features and the use of the non-linear classifier did not lead to the differentiability between MDD and HC. Our results support the notion that MDD classification on this combination of features and classifiers is unfeasible.
Wearable data from subjects playing Super Mario, sitting university exams, or performing physical exercise help detect acute mood episodes via self-supervised learning
Nov 07, 2023



Abstract:Personal sensing, leveraging data passively and near-continuously collected with wearables from patients in their ecological environment, is a promising paradigm to monitor mood disorders (MDs), a major determinant of worldwide disease burden. However, collecting and annotating wearable data is very resource-intensive. Studies of this kind can thus typically afford to recruit only a couple dozens of patients. This constitutes one of the major obstacles to applying modern supervised machine learning techniques to MDs detection. In this paper, we overcome this data bottleneck and advance the detection of MDs acute episode vs stable state from wearables data on the back of recent advances in self-supervised learning (SSL). This leverages unlabelled data to learn representations during pre-training, subsequently exploited for a supervised task. First, we collected open-access datasets recording with an Empatica E4 spanning different, unrelated to MD monitoring, personal sensing tasks -- from emotion recognition in Super Mario players to stress detection in undergraduates -- and devised a pre-processing pipeline performing on-/off-body detection, sleep-wake detection, segmentation, and (optionally) feature extraction. With 161 E4-recorded subjects, we introduce E4SelfLearning, the largest to date open access collection, and its pre-processing pipeline. Second, we show that SSL confidently outperforms fully-supervised pipelines using either our novel E4-tailored Transformer architecture (E4mer) or classical baseline XGBoost: 81.23% against 75.35% (E4mer) and 72.02% (XGBoost) correctly classified recording segments from 64 (half acute, half stable) patients. Lastly, we illustrate that SSL performance is strongly associated with the specific surrogate task employed for pre-training as well as with unlabelled data availability.
Analysis of an Automated Machine Learning Approach in Brain Predictive Modelling: A data-driven approach to Predict Brain Age from Cortical Anatomical Measures
Oct 08, 2019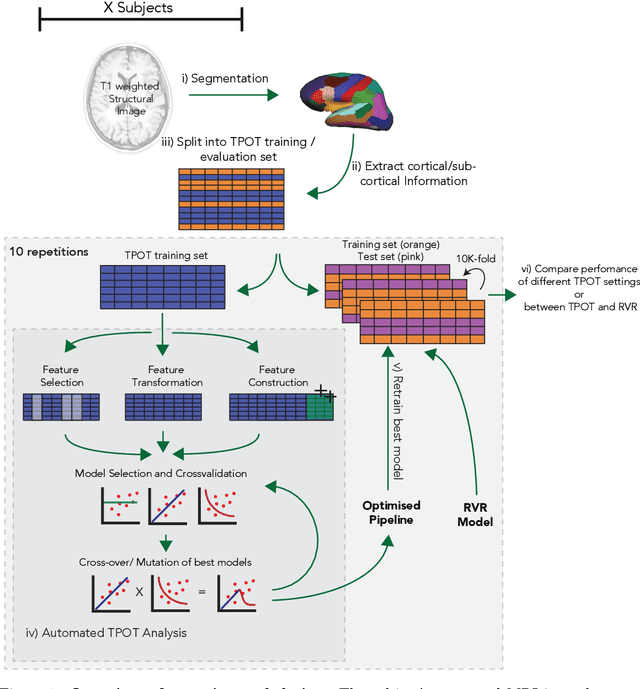


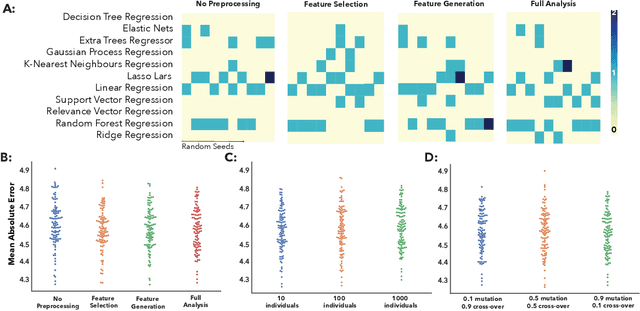
Abstract:The use of machine learning (ML) algorithms has significantly increased in neuroscience. However, from the vast extent of possible ML algorithms, which one is the optimal model to predict the target variable? What are the hyperparameters for such a model? Given the plethora of possible answers to these questions, in the last years, automated machine learning (autoML) has been gaining attention. Here, we apply an autoML library called TPOT which uses a tree-based representation of machine learning pipelines and conducts a genetic-programming based approach to find the model and its hyperparameters that more closely predicts the subject's true age. To explore autoML and evaluate its efficacy within neuroimaging datasets, we chose a problem that has been the focus of previous extensive study: brain age prediction. Without any prior knowledge, TPOT was able to scan through the model space and create pipelines that outperformed the state-of-the-art accuracy for Freesurfer-based models using only thickness and volume information for anatomical structure. In particular, we compared the performance of TPOT (mean accuracy error (MAE): $4.612 \pm .124$ years) and a Relevance Vector Regression (MAE $5.474 \pm .140$ years). TPOT also suggested interesting combinations of models that do not match the current most used models for brain prediction but generalise well to unseen data. AutoML showed promising results as a data-driven approach to find optimal models for neuroimaging applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge