George Ioannakis
When Slots Compete: Slot Merging in Object-Centric Learning
Mar 11, 2026Abstract:Slot-based object-centric learning represents an image as a set of latent slots with a decoder that combines them into an image or features. The decoder specifies how slots are combined into an output, but the slot set is typically fixed: the number of slots is chosen upfront and slots are only refined. This can lead to multiple slots competing for overlapping regions of the same entity rather than focusing on distinct regions. We introduce slot merging: a drop-in, lightweight operation on the slot set that merges overlapping slots during training. We quantify overlap with a Soft-IoU score between slot-attention maps and combine selected pairs via a barycentric update that preserves gradient flow. Merging follows a fixed policy, with the decision threshold inferred from overlap statistics, requiring no additional learnable modules. Integrated into the established feature-reconstruction pipeline of DINOSAUR, the proposed method improves object factorization and mask quality, surpassing other adaptive methods in object discovery and segmentation benchmarks.
ART3mis: Ray-Based Textual Annotation on 3D Cultural Objects
Feb 13, 2026Abstract:Beyond simplistic 3D visualisations, archaeologists, as well as cultural heritage experts and practitioners, need applications with advanced functionalities. Such as the annotation and attachment of metadata onto particular regions of the 3D digital objects. Various approaches have been presented to tackle this challenge, most of which achieve excellent results in the domain of their application. However, they are often confined to that specific domain and particular problem. In this paper, we present ART3mis - a general-purpose, user-friendly, interactive textual annotation tool for 3D objects. Primarily attuned to aid cultural heritage conservators, restorers and curators with no technical skills in 3D imaging and graphics, the tool allows for the easy handling, segmenting and annotating of 3D digital replicas of artefacts. ART3mis applies a user-driven, direct-on-surface approach. It can handle detailed 3D cultural objects in real-time and store textual annotations for multiple complex regions in JSON data format.
GARLIC: GAussian Representation LearnIng for spaCe partitioning
May 30, 2025Abstract:We introduce GARLIC (GAussian Representation LearnIng for spaCe partitioning), a novel indexing structure based on \(N\)-dimensional Gaussians for efficiently learning high-dimensional vector spaces. Our approach is inspired from Gaussian splatting techniques, typically used in 3D rendering, which we adapt for high-dimensional search and classification. We optimize Gaussian parameters using information-theoretic objectives that balance coverage, assignment confidence, and structural and semantic consistency. A key contribution is to progressively refine the representation through split and clone operations, handling hundreds of dimensions, thus handling varying data densities. GARLIC offers the fast building times of traditional space partitioning methods (e.g., under \(\sim5\) min build time for SIFT1M) while achieving \(\sim50\%\) Recall10@10 in low-candidate regimes. Experimental results on standard benchmarks demonstrate our method's consistency in (a) \(k\)-NN retrieval, outperforming methods, such as Faiss-IVF, in fast-recall by using about half their probes for the same Recall10@10 in Fashion-MNIST, and (b) in classification tasks, beating by \(\sim15\%\) accuracy other majority voting methods. Further, we show strong generalization capabilities, maintaining high accuracy even with downsampled training data: using just \(1\%\) of the training data returns \(\sim 45\%\) Recall@1, thus making GARLIC quite powerful for applications requiring both speed and accuracy.
Semantic Image Segmentation with Deep Learning for Vine Leaf Phenotyping
Oct 24, 2022



Abstract:Plant phenotyping refers to a quantitative description of the plants properties, however in image-based phenotyping analysis, our focus is primarily on the plants anatomical, ontogenetical and physiological properties.This technique reinforced by the success of Deep Learning in the field of image based analysis is applicable to a wide range of research areas making high-throughput screens of plants possible, reducing the time and effort needed for phenotypic characterization.In this study, we use Deep Learning methods (supervised and unsupervised learning based approaches) to semantically segment grapevine leaves images in order to develop an automated object detection (through segmentation) system for leaf phenotyping which will yield information regarding their structure and function.In these directions we studied several deep learning approaches with promising results as well as we reported some future challenging tasks in the area of precision agriculture.Our work contributes to plant lifecycle monitoring through which dynamic traits such as growth and development can be captured and quantified, targeted intervention and selective application of agrochemicals and grapevine variety identification which are key prerequisites in sustainable agriculture.
Pulmonary embolism identification in computerized tomography pulmonary angiography scans with deep learning technologies in COVID-19 patients
May 28, 2021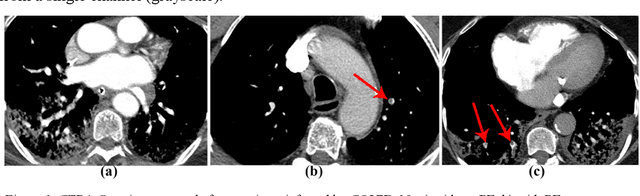
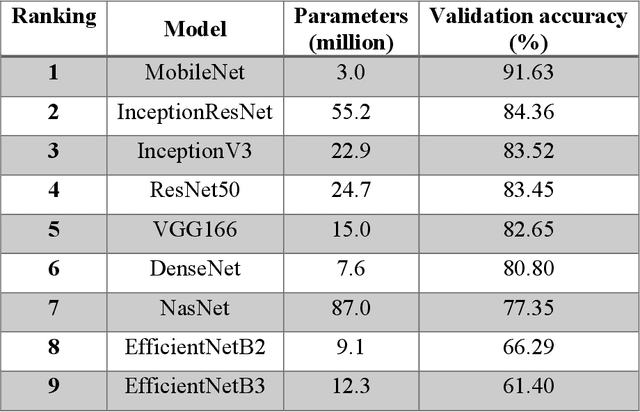
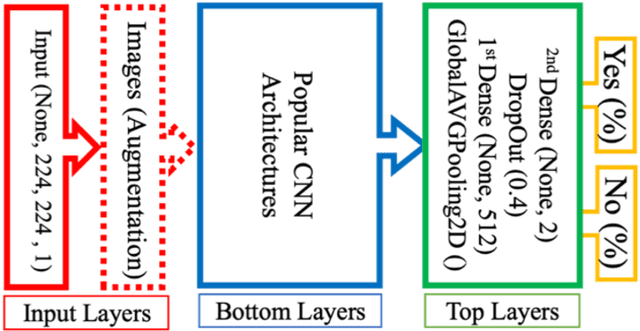
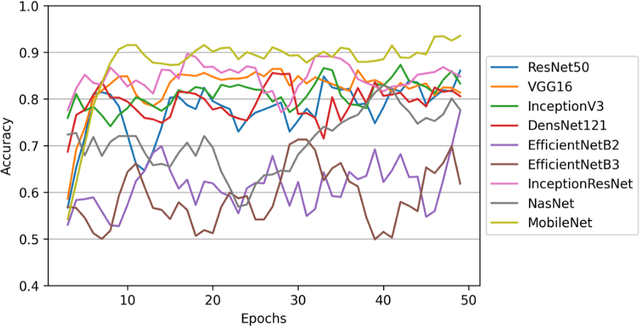
Abstract:The main objective of this work is to utilize state-of-the-art deep learning approaches for the identification of pulmonary embolism in CTPA-Scans for COVID-19 patients, provide an initial assessment of their performance and, ultimately, provide a fast-track prototype solution (system). We adopted and assessed some of the most popular convolutional neural network architectures through transfer learning approaches, to strive to combine good model accuracy with fast training. Additionally, we exploited one of the most popular one-stage object detection models for the localization (through object detection) of the pulmonary embolism regions-of-interests. The models of both approaches are trained on an original CTPA-Scan dataset, where we annotated of 673 CTPA-Scan images with 1,465 bounding boxes in total, highlighting pulmonary embolism regions-of-interests. We provide a brief assessment of some state-of-the-art image classification models by achieving validation accuracies of 91% in pulmonary embolism classification. Additionally, we achieved a precision of about 68% on average in the object detection model for the pulmonary embolism localization under 50% IoU threshold. For both approaches, we provide the entire training pipelines for future studies (step by step processes through source code). In this study, we present some of the most accurate and fast deep learning models for pulmonary embolism identification in CTPA-Scans images, through classification and localization (object detection) approaches for patients infected by COVID-19. We provide a fast-track solution (system) for the research community of the area, which combines both classification and object detection models for improving the precision of identifying pulmonary embolisms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge