Farzan Moodi
Attention Xception UNet (AXUNet): A Novel Combination of CNN and Self-Attention for Brain Tumor Segmentation
Mar 26, 2025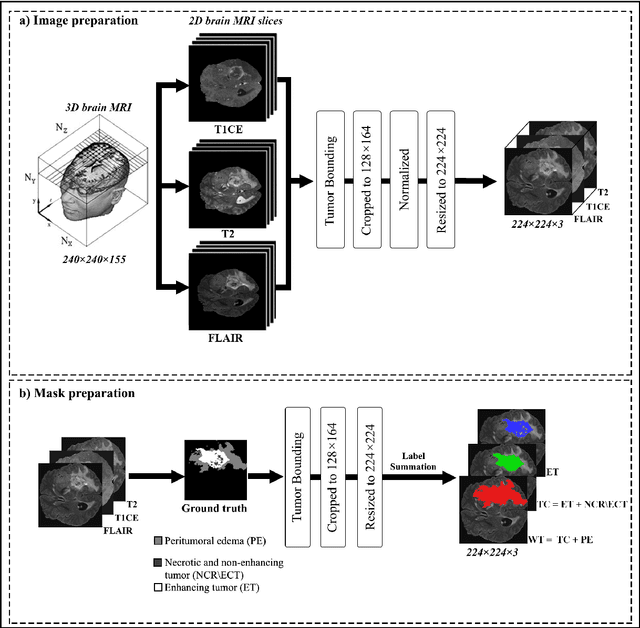


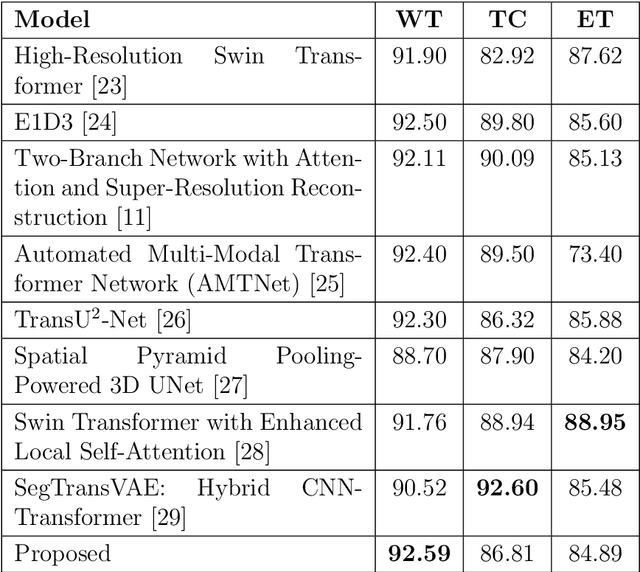
Abstract:Accurate segmentation of glioma brain tumors is crucial for diagnosis and treatment planning. Deep learning techniques offer promising solutions, but optimal model architectures remain under investigation. We used the BraTS 2021 dataset, selecting T1 with contrast enhancement (T1CE), T2, and Fluid-Attenuated Inversion Recovery (FLAIR) sequences for model development. The proposed Attention Xception UNet (AXUNet) architecture integrates an Xception backbone with dot-product self-attention modules, inspired by state-of-the-art (SOTA) large language models such as Google Bard and OpenAI ChatGPT, within a UNet-shaped model. We compared AXUNet with SOTA models. Comparative evaluation on the test set demonstrated improved results over baseline models. Inception-UNet and Xception-UNet achieved mean Dice scores of 90.88 and 93.24, respectively. Attention ResUNet (AResUNet) attained a mean Dice score of 92.80, with the highest score of 84.92 for enhancing tumor (ET) among all models. Attention Gate UNet (AGUNet) yielded a mean Dice score of 90.38. AXUNet outperformed all models with a mean Dice score of 93.73. It demonstrated superior Dice scores across whole tumor (WT) and tumor core (TC) regions, achieving 92.59 for WT, 86.81 for TC, and 84.89 for ET. The integration of the Xception backbone and dot-product self-attention mechanisms in AXUNet showcases enhanced performance in capturing spatial and contextual information. The findings underscore the potential utility of AXUNet in facilitating precise tumor delineation.
The 2024 Brain Tumor Segmentation (BraTS) Challenge: Glioma Segmentation on Post-treatment MRI
May 28, 2024
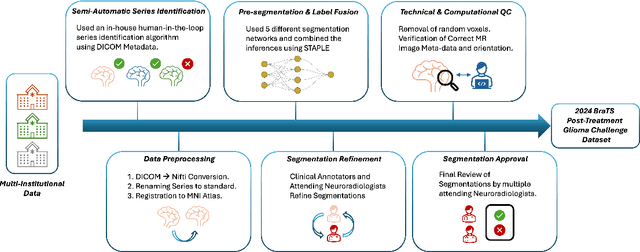
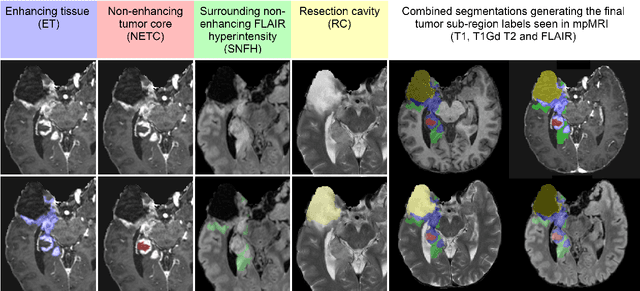
Abstract:Gliomas are the most common malignant primary brain tumors in adults and one of the deadliest types of cancer. There are many challenges in treatment and monitoring due to the genetic diversity and high intrinsic heterogeneity in appearance, shape, histology, and treatment response. Treatments include surgery, radiation, and systemic therapies, with magnetic resonance imaging (MRI) playing a key role in treatment planning and post-treatment longitudinal assessment. The 2024 Brain Tumor Segmentation (BraTS) challenge on post-treatment glioma MRI will provide a community standard and benchmark for state-of-the-art automated segmentation models based on the largest expert-annotated post-treatment glioma MRI dataset. Challenge competitors will develop automated segmentation models to predict four distinct tumor sub-regions consisting of enhancing tissue (ET), surrounding non-enhancing T2/fluid-attenuated inversion recovery (FLAIR) hyperintensity (SNFH), non-enhancing tumor core (NETC), and resection cavity (RC). Models will be evaluated on separate validation and test datasets using standardized performance metrics utilized across the BraTS 2024 cluster of challenges, including lesion-wise Dice Similarity Coefficient and Hausdorff Distance. Models developed during this challenge will advance the field of automated MRI segmentation and contribute to their integration into clinical practice, ultimately enhancing patient care.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge