Fabio S. Ferreira
Identifying latent disease factors differently expressed in patient subgroups using group factor analysis
Oct 10, 2024



Abstract:In this study, we propose a novel approach to uncover subgroup-specific and subgroup-common latent factors addressing the challenges posed by the heterogeneity of neurological and mental disorders, which hinder disease understanding, treatment development, and outcome prediction. The proposed approach, sparse Group Factor Analysis (GFA) with regularised horseshoe priors, was implemented with probabilistic programming and can uncover associations (or latent factors) among multiple data modalities differentially expressed in sample subgroups. Synthetic data experiments showed the robustness of our sparse GFA by correctly inferring latent factors and model parameters. When applied to the Genetic Frontotemporal Dementia Initiative (GENFI) dataset, which comprises patients with frontotemporal dementia (FTD) with genetically defined subgroups, the sparse GFA identified latent disease factors differentially expressed across the subgroups, distinguishing between "subgroup-specific" latent factors within homogeneous groups and "subgroup common" latent factors shared across subgroups. The latent disease factors captured associations between brain structure and non-imaging variables (i.e., questionnaires assessing behaviour and disease severity) across the different genetic subgroups, offering insights into disease profiles. Importantly, two latent factors were more pronounced in the two more homogeneous FTD patient subgroups (progranulin (GRN) and microtubule-associated protein tau (MAPT) mutation), showcasing the method's ability to reveal subgroup-specific characteristics. These findings underscore the potential of sparse GFA for integrating multiple data modalities and identifying interpretable latent disease factors that can improve the characterization and stratification of patients with neurological and mental health disorders.
A hierarchical Bayesian model to find brain-behaviour associations in incomplete data sets
Mar 11, 2021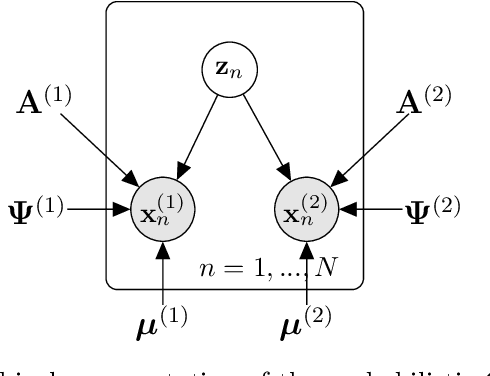

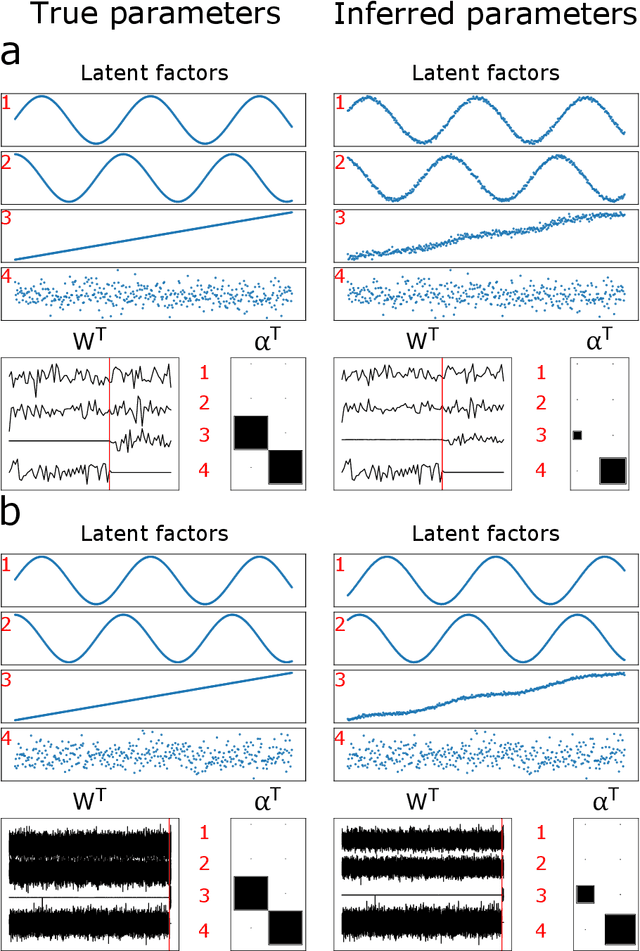

Abstract:Canonical Correlation Analysis (CCA) and its regularised versions have been widely used in the neuroimaging community to uncover multivariate associations between two data modalities (e.g., brain imaging and behaviour). However, these methods have inherent limitations: (1) statistical inferences about the associations are often not robust; (2) the associations within each data modality are not modelled; (3) missing values need to be imputed or removed. Group Factor Analysis (GFA) is a hierarchical model that addresses the first two limitations by providing Bayesian inference and modelling modality-specific associations. Here, we propose an extension of GFA that handles missing data, and highlight that GFA can be used as a predictive model. We applied GFA to synthetic and real data consisting of brain connectivity and non-imaging measures from the Human Connectome Project (HCP). In synthetic data, GFA uncovered the underlying shared and specific factors and predicted correctly the non-observed data modalities in complete and incomplete data sets. In the HCP data, we identified four relevant shared factors, capturing associations between mood, alcohol and drug use, cognition, demographics and psychopathological measures and the default mode, frontoparietal control, dorsal and ventral networks and insula, as well as two factors describing associations within brain connectivity. In addition, GFA predicted a set of non-imaging measures from brain connectivity. These findings were consistent in complete and incomplete data sets, and replicated previous findings in the literature. GFA is a promising tool that can be used to uncover associations between and within multiple data modalities in benchmark datasets (such as, HCP), and easily extended to more complex models to solve more challenging tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge