Ezequiel Geremia
3D-Morphomics, Morphological Features on CT scans for lung nodule malignancy diagnosis
Jul 27, 2022
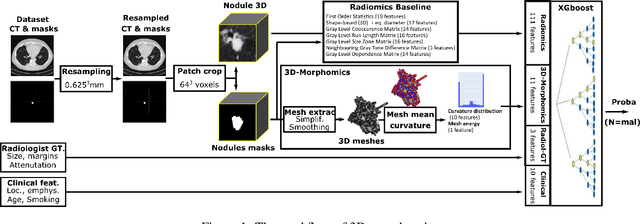
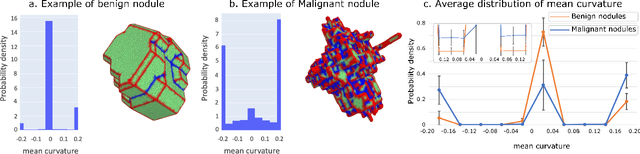
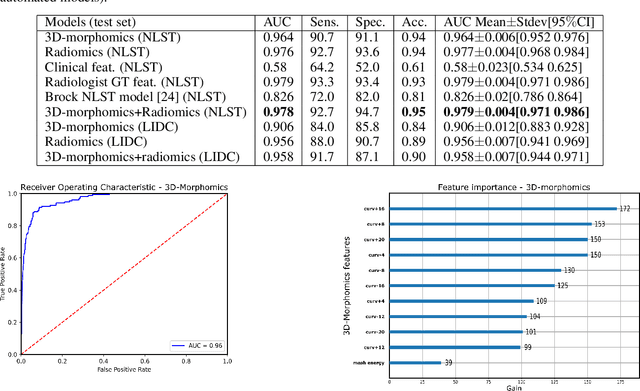
Abstract:Pathologies systematically induce morphological changes, thus providing a major but yet insufficiently quantified source of observables for diagnosis. The study develops a predictive model of the pathological states based on morphological features (3D-morphomics) on Computed Tomography (CT) volumes. A complete workflow for mesh extraction and simplification of an organ's surface is developed, and coupled with an automatic extraction of morphological features given by the distribution of mean curvature and mesh energy. An XGBoost supervised classifier is then trained and tested on the 3D-morphomics to predict the pathological states. This framework is applied to the prediction of the malignancy of lung's nodules. On a subset of NLST database with malignancy confirmed biopsy, using 3D-morphomics only, the classification model of lung nodules into malignant vs. benign achieves 0.964 of AUC. Three other sets of classical features are trained and tested, (1) clinical relevant features gives an AUC of 0.58, (2) 111 radiomics gives an AUC of 0.976, (3) radiologist ground truth (GT) containing the nodule size, attenuation and spiculation qualitative annotations gives an AUC of 0.979. We also test the Brock model and obtain an AUC of 0.826. Combining 3D-morphomics and radiomics features achieves state-of-the-art results with an AUC of 0.978 where the 3D-morphomics have some of the highest predictive powers. As a validation on a public independent cohort, models are applied to the LIDC dataset, the 3D-morphomics achieves an AUC of 0.906 and the 3D-morphomics+radiomics achieves an AUC of 0.958, which ranks second in the challenge among deep models. It establishes the curvature distributions as efficient features for predicting lung nodule malignancy and a new method that can be applied directly to arbitrary computer aided diagnosis task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge