David M. Groppe
Node-Centric Graph Learning from Data for Brain State Identification
Nov 04, 2020
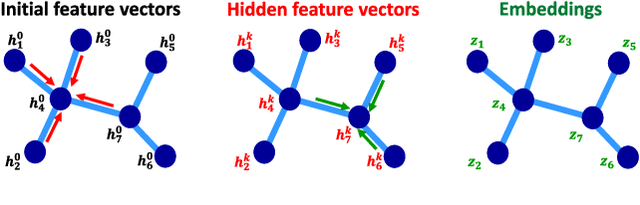
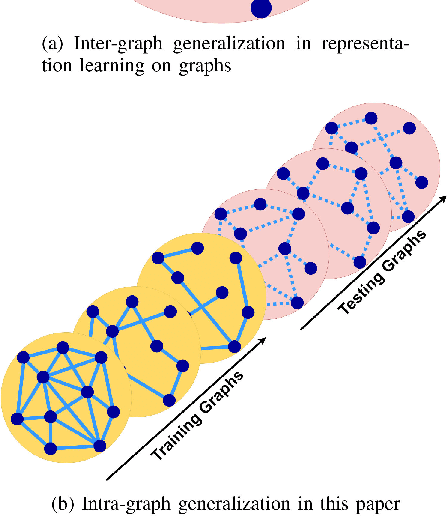
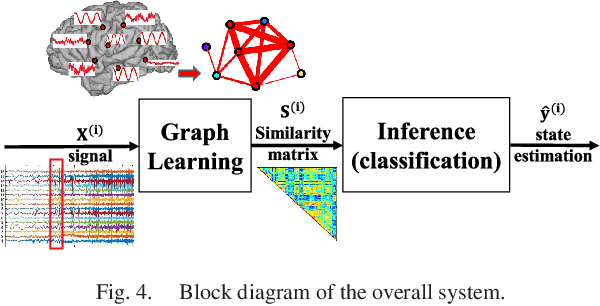
Abstract:Data-driven graph learning models a network by determining the strength of connections between its nodes. The data refers to a graph signal which associates a value with each graph node. Existing graph learning methods either use simplified models for the graph signal, or they are prohibitively expensive in terms of computational and memory requirements. This is particularly true when the number of nodes is high or there are temporal changes in the network. In order to consider richer models with a reasonable computational tractability, we introduce a graph learning method based on representation learning on graphs. Representation learning generates an embedding for each graph node, taking the information from neighbouring nodes into account. Our graph learning method further modifies the embeddings to compute the graph similarity matrix. In this work, graph learning is used to examine brain networks for brain state identification. We infer time-varying brain graphs from an extensive dataset of intracranial electroencephalographic (iEEG) signals from ten patients. We then apply the graphs as input to a classifier to distinguish seizure vs. non-seizure brain states. Using the binary classification metric of area under the receiver operating characteristic curve (AUC), this approach yields an average of 9.13 percent improvement when compared to two widely used brain network modeling methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge