David Bendahan
CNN-based fully automatic wrist cartilage volume quantification in MR Image
Jun 22, 2022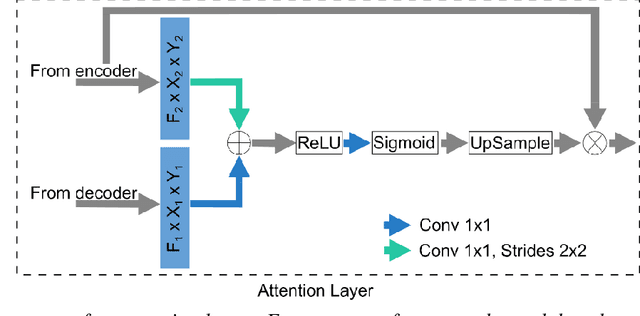
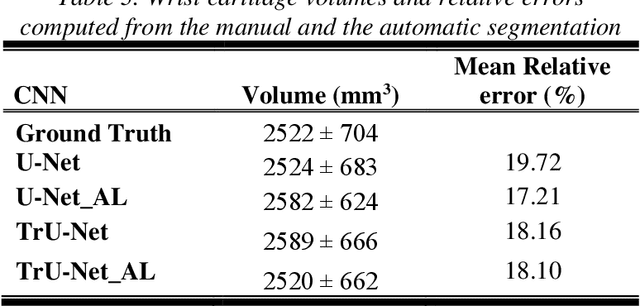

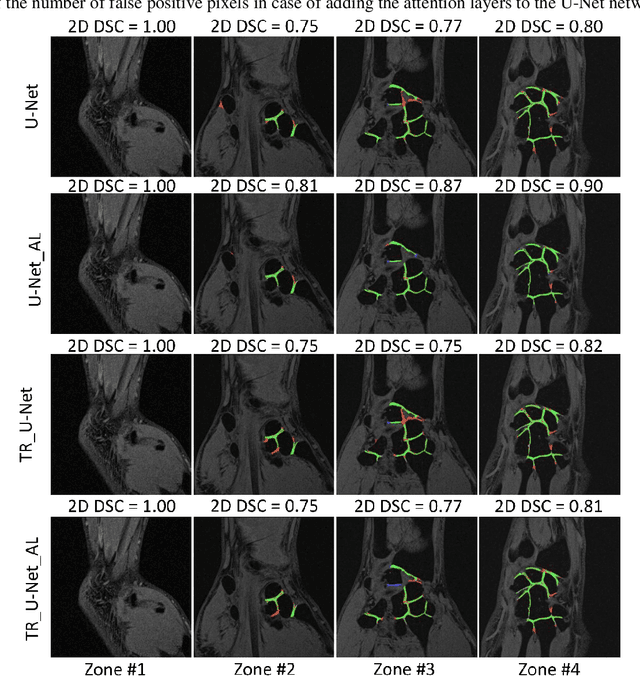
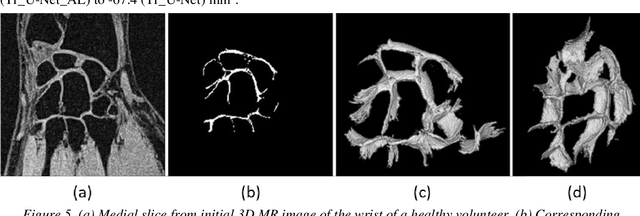
Abstract:Detection of cartilage loss is crucial for the diagnosis of osteo- and rheumatoid arthritis. A large number of automatic segmentation tools have been reported so far for cartilage assessment in magnetic resonance images of large joints. As compared to knee or hip, wrist cartilage has a more complex structure so that automatic tools developed for large joints are not expected to be operational for wrist cartilage segmentation. In that respect, a fully automatic wrist cartilage segmentation method would be of high clinical interest. We assessed the performance of four optimized variants of the U-Net architecture with truncation of its depth and addition of attention layers (U-Net_AL). The corresponding results were compared to those from a patch-based convolutional neural network (CNN) we previously designed. The segmentation quality was assessed on the basis of a comparative analysis with manual segmentation using several morphological (2D DSC, 3D DSC, precision) and a volumetric metrics. The four networks outperformed the patch-based CNN in terms of segmentation homogeneity and quality. The median 3D DSC value computed with the U-Net_AL (0.817) was significantly larger than the corresponding DSC values computed with the other networks. In addition, the U-Net_AL CNN provided the lowest mean volume error (17%) and the highest Pearson correlation coefficient (0.765) with respect to the ground truth. Of interest, the reproducibility computed from using U-Net_AL was larger than the reproducibility of the manual segmentation. U-net convolutional neural network with additional attention layers provides the best wrist cartilage segmentation performance. In order to be used in clinical conditions, the trained network can be fine-tuned on a dataset representing a group of specific patients. The error of cartilage volume measurement should be assessed independently using a non-MRI method.
Automatic Segmentation of Muscle Tissue and Inter-muscular Fat in Thigh and Calf MRI Images
Oct 01, 2019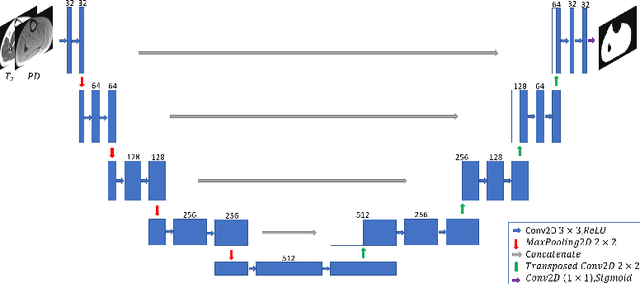


Abstract:Magnetic resonance imaging (MRI) of thigh and calf muscles is one of the most effective techniques for estimating fat infiltration into muscular dystrophies. The infiltration of adipose tissue into the diseased muscle region varies in its severity across, and within, patients. In order to efficiently quantify the infiltration of fat, accurate segmentation of muscle and fat is needed. An estimation of the amount of infiltrated fat is typically done visually by experts. Several algorithmic solutions have been proposed for automatic segmentation. While these methods may work well in mild cases, they struggle in moderate and severe cases due to the high variability in the intensity of infiltration, and the tissue's heterogeneous nature. To address these challenges, we propose a deep-learning approach, producing robust results with high Dice Similarity Coefficient (DSC) of 0.964, 0.917 and 0.933 for muscle-region, healthy muscle and inter-muscular adipose tissue (IMAT) segmentation, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge