Daniele Santoni
TFBS-Finder: Deep Learning-based Model with DNABERT and Convolutional Networks to Predict Transcription Factor Binding Sites
Feb 03, 2025



Abstract:Transcription factors are proteins that regulate the expression of genes by binding to specific genomic regions known as Transcription Factor Binding Sites (TFBSs), typically located in the promoter regions of those genes. Accurate prediction of these binding sites is essential for understanding the complex gene regulatory networks underlying various cellular functions. In this regard, many deep learning models have been developed for such prediction, but there is still scope of improvement. In this work, we have developed a deep learning model which uses pre-trained DNABERT, a Convolutional Neural Network (CNN) module, a Modified Convolutional Block Attention Module (MCBAM), a Multi-Scale Convolutions with Attention (MSCA) module and an output module. The pre-trained DNABERT is used for sequence embedding, thereby capturing the long-term dependencies in the DNA sequences while the CNN, MCBAM and MSCA modules are useful in extracting higher-order local features. TFBS-Finder is trained and tested on 165 ENCODE ChIP-seq datasets. We have also performed ablation studies as well as cross-cell line validations and comparisons with other models. The experimental results show the superiority of the proposed method in predicting TFBSs compared to the existing methodologies. The codes and the relevant datasets are publicly available at https://github.com/NimishaGhosh/TFBS-Finder/.
A Review on the Applications of Transformer-based language models for Nucleotide Sequence Analysis
Dec 10, 2024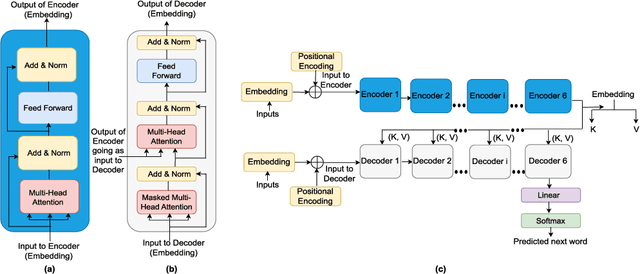

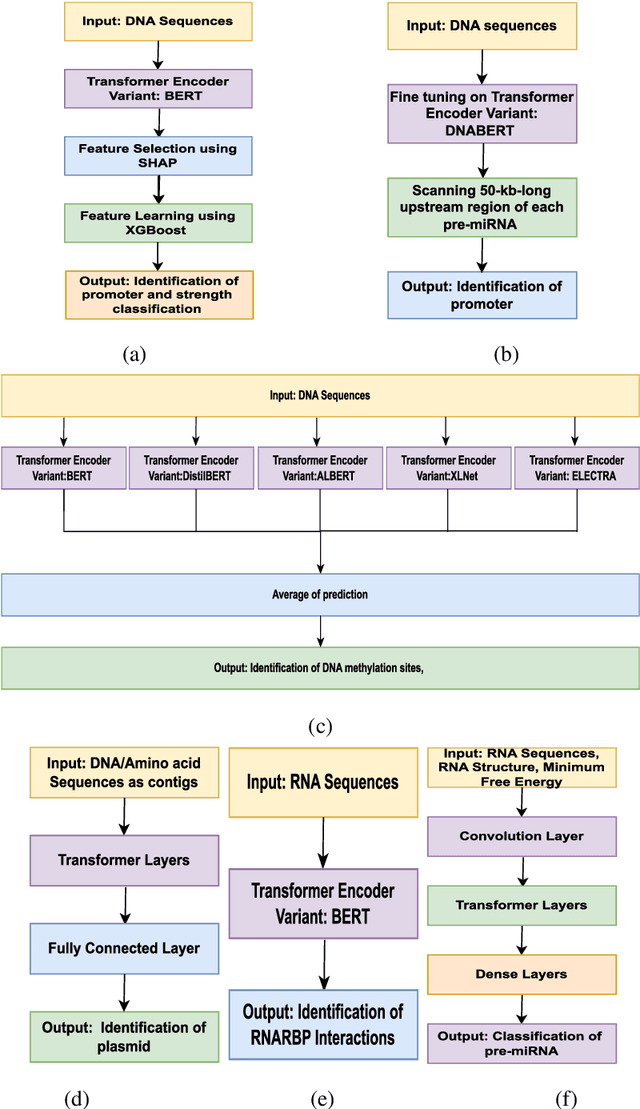
Abstract:In recent times, Transformer-based language models are making quite an impact in the field of natural language processing. As relevant parallels can be drawn between biological sequences and natural languages, the models used in NLP can be easily extended and adapted for various applications in bioinformatics. In this regard, this paper introduces the major developments of Transformer-based models in the recent past in the context of nucleotide sequences. We have reviewed and analysed a large number of application-based papers on this subject, giving evidence of the main characterizing features and to different approaches that may be adopted to customize such powerful computational machines. We have also provided a structured description of the functioning of Transformers, that may enable even first time users to grab the essence of such complex architectures. We believe this review will help the scientific community in understanding the various applications of Transformer-based language models to nucleotide sequences. This work will motivate the readers to build on these methodologies to tackle also various other problems in the field of bioinformatics.
Predicting Transcription Factor Binding Sites using Transformer based Capsule Network
Oct 23, 2023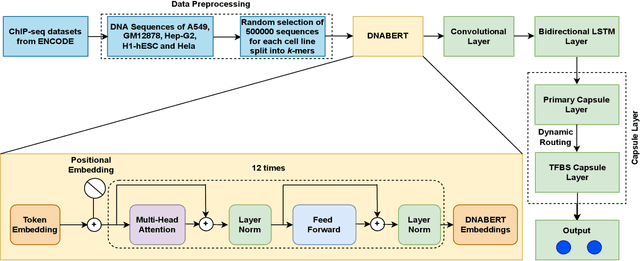
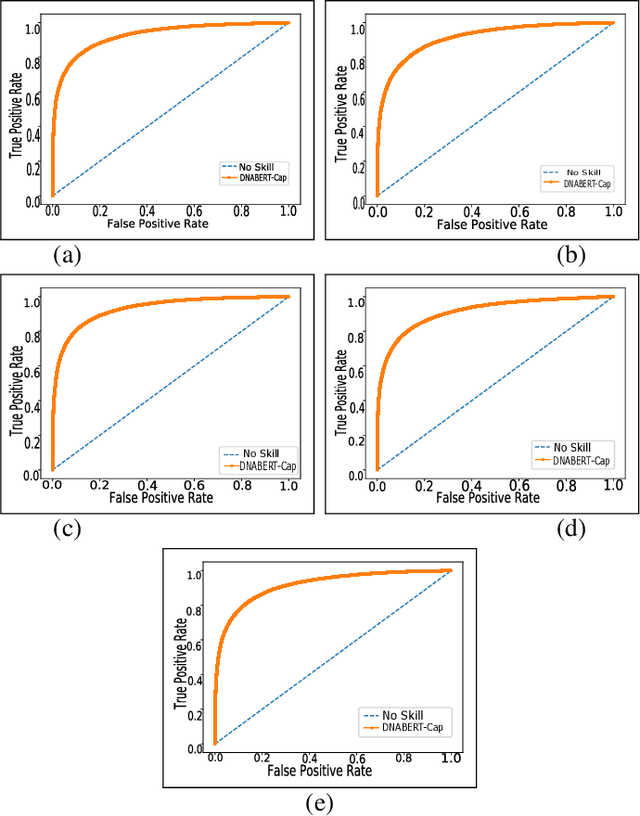
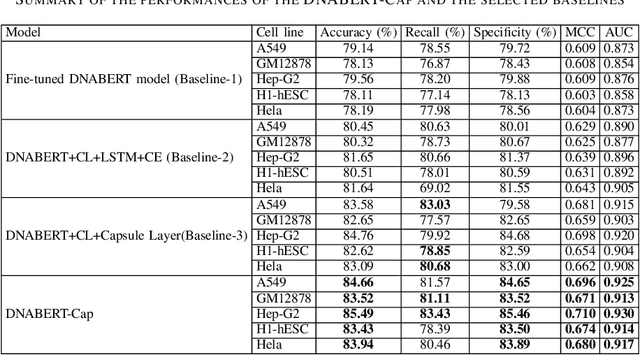
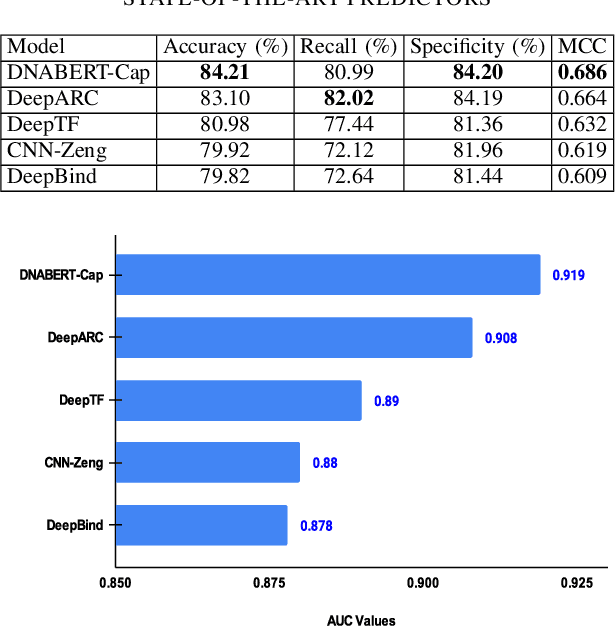
Abstract:Prediction of binding sites for transcription factors is important to understand how they regulate gene expression and how this regulation can be modulated for therapeutic purposes. Although in the past few years there are significant works addressing this issue, there is still space for improvement. In this regard, a transformer based capsule network viz. DNABERT-Cap is proposed in this work to predict transcription factor binding sites mining ChIP-seq datasets. DNABERT-Cap is a bidirectional encoder pre-trained with large number of genomic DNA sequences, empowered with a capsule layer responsible for the final prediction. The proposed model builds a predictor for transcription factor binding sites using the joint optimisation of features encompassing both bidirectional encoder and capsule layer, along with convolutional and bidirectional long-short term memory layers. To evaluate the efficiency of the proposed approach, we use a benchmark ChIP-seq datasets of five cell lines viz. A549, GM12878, Hep-G2, H1-hESC and Hela, available in the ENCODE repository. The results show that the average area under the receiver operating characteristic curve score exceeds 0.91 for all such five cell lines. DNABERT-Cap is also compared with existing state-of-the-art deep learning based predictors viz. DeepARC, DeepTF, CNN-Zeng and DeepBind, and is seen to outperform them.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge