Daniel Tward
Individualized multi-horizon MRI trajectory prediction for Alzheimer's Disease
Aug 04, 2024



Abstract:Neurodegeneration as measured through magnetic resonance imaging (MRI) is recognized as a potential biomarker for diagnosing Alzheimer's disease (AD), but is generally considered less specific than amyloid or tau based biomarkers. Due to a large amount of variability in brain anatomy between different individuals, we hypothesize that leveraging MRI time series can help improve specificity, by treating each patient as their own baseline. Here we turn to conditional variational autoencoders to generate individualized MRI predictions given the subject's age, disease status and one previous scan. Using serial imaging data from the Alzheimer's Disease Neuroimaging Initiative, we train a novel architecture to build a latent space distribution which can be sampled from to generate future predictions of changing anatomy. This enables us to extrapolate beyond the dataset and predict MRIs up to 10 years. We evaluated the model on a held-out set from ADNI and an independent dataset (from Open Access Series of Imaging Studies). By comparing to several alternatives, we show that our model produces more individualized images with higher resolution. Further, if an individual already has a follow-up MRI, we demonstrate a usage of our model to compute a likelihood ratio classifier for disease status. In practice, the model may be able to assist in early diagnosis of AD and provide a counterfactual baseline trajectory for treatment effect estimation. Furthermore, it generates a synthetic dataset that can potentially be used for downstream tasks such as anomaly detection and classification.
Hidden Markov Modeling for Maximum Likelihood Neuron Reconstruction
Jun 04, 2021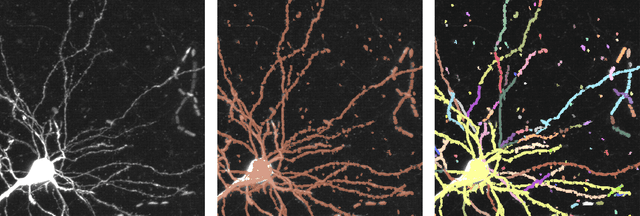

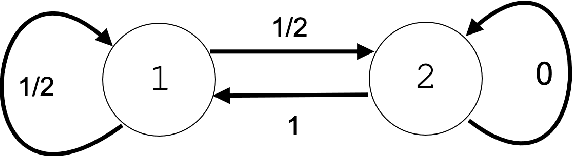

Abstract:Recent advances in brain clearing and imaging have made it possible to image entire mammalian brains at sub-micron resolution. These images offer the potential to assemble brain-wide atlases of projection neuron morphology, but manual neuron reconstruction remains a bottleneck. Here we present a method inspired by hidden Markov modeling and appearance modeling of fluorescent neuron images that can automatically trace neuronal processes. Our method leverages dynamic programming to scale to terabyte sized image data and can be applied to images with one or more neurons. We applied our algorithm to the output of image segmentation models where false negatives severed neuronal processes, and showed that it can follow axons in the presence of noise or nearby neurons. Our method has the potential to be integrated into a semi or fully automated reconstruction pipeline. Additionally, it creates a framework through which users can intervene with hard constraints to, for example, rule out certain reconstructions, or assign axons to particular cell bodies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge