Daniel Scholz
VariViT: A Vision Transformer for Variable Image Sizes
Feb 16, 2026Abstract:Vision Transformers (ViTs) have emerged as the state-of-the-art architecture in representation learning, leveraging self-attention mechanisms to excel in various tasks. ViTs split images into fixed-size patches, constraining them to a predefined size and necessitating pre-processing steps like resizing, padding, or cropping. This poses challenges in medical imaging, particularly with irregularly shaped structures like tumors. A fixed bounding box crop size produces input images with highly variable foreground-to-background ratios. Resizing medical images can degrade information and introduce artefacts, impacting diagnosis. Hence, tailoring variable-sized crops to regions of interest can enhance feature representation capabilities. Moreover, large images are computationally expensive, and smaller sizes risk information loss, presenting a computation-accuracy tradeoff. We propose VariViT, an improved ViT model crafted to handle variable image sizes while maintaining a consistent patch size. VariViT employs a novel positional embedding resizing scheme for a variable number of patches. We also implement a new batching strategy within VariViT to reduce computational complexity, resulting in faster training and inference times. In our evaluations on two 3D brain MRI datasets, VariViT surpasses vanilla ViTs and ResNet in glioma genotype prediction and brain tumor classification. It achieves F1-scores of 75.5% and 76.3%, respectively, learning more discriminative features. Our proposed batching strategy reduces computation time by up to 30% compared to conventional architectures. These findings underscore the efficacy of VariViT in image representation learning. Our code can be found here: https://github.com/Aswathi-Varma/varivit
All Sizes Matter: Improving Volumetric Brain Segmentation on Small Lesions
Oct 04, 2023



Abstract:Brain metastases (BMs) are the most frequently occurring brain tumors. The treatment of patients having multiple BMs with stereo tactic radiosurgery necessitates accurate localization of the metastases. Neural networks can assist in this time-consuming and costly task that is typically performed by human experts. Particularly challenging is the detection of small lesions since they are often underrepresented in exist ing approaches. Yet, lesion detection is equally important for all sizes. In this work, we develop an ensemble of neural networks explicitly fo cused on detecting and segmenting small BMs. To accomplish this task, we trained several neural networks focusing on individual aspects of the BM segmentation problem: We use blob loss that specifically addresses the imbalance of lesion instances in terms of size and texture and is, therefore, not biased towards larger lesions. In addition, a model using a subtraction sequence between the T1 and T1 contrast-enhanced sequence focuses on low-contrast lesions. Furthermore, we train additional models only on small lesions. Our experiments demonstrate the utility of the ad ditional blob loss and the subtraction sequence. However, including the specialized small lesion models in the ensemble deteriorates segmentation results. We also find domain-knowledge-inspired postprocessing steps to drastically increase our performance in most experiments. Our approach enables us to submit a competitive challenge entry to the ASNR-MICCAI BraTS Brain Metastasis Challenge 2023.
Metrics to Quantify Global Consistency in Synthetic Medical Images
Aug 01, 2023


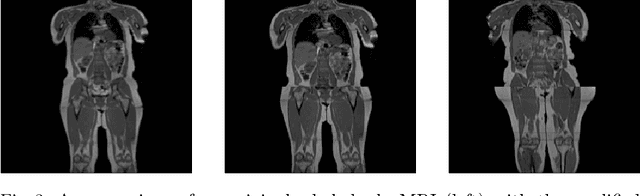
Abstract:Image synthesis is increasingly being adopted in medical image processing, for example for data augmentation or inter-modality image translation. In these critical applications, the generated images must fulfill a high standard of biological correctness. A particular requirement for these images is global consistency, i.e an image being overall coherent and structured so that all parts of the image fit together in a realistic and meaningful way. Yet, established image quality metrics do not explicitly quantify this property of synthetic images. In this work, we introduce two metrics that can measure the global consistency of synthetic images on a per-image basis. To measure the global consistency, we presume that a realistic image exhibits consistent properties, e.g., a person's body fat in a whole-body MRI, throughout the depicted object or scene. Hence, we quantify global consistency by predicting and comparing explicit attributes of images on patches using supervised trained neural networks. Next, we adapt this strategy to an unlabeled setting by measuring the similarity of implicit image features predicted by a self-supervised trained network. Our results demonstrate that predicting explicit attributes of synthetic images on patches can distinguish globally consistent from inconsistent images. Implicit representations of images are less sensitive to assess global consistency but are still serviceable when labeled data is unavailable. Compared to established metrics, such as the FID, our method can explicitly measure global consistency on a per-image basis, enabling a dedicated analysis of the biological plausibility of single synthetic images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge