Daniel G. Chen
Celcomen: spatial causal disentanglement for single-cell and tissue perturbation modeling
Sep 09, 2024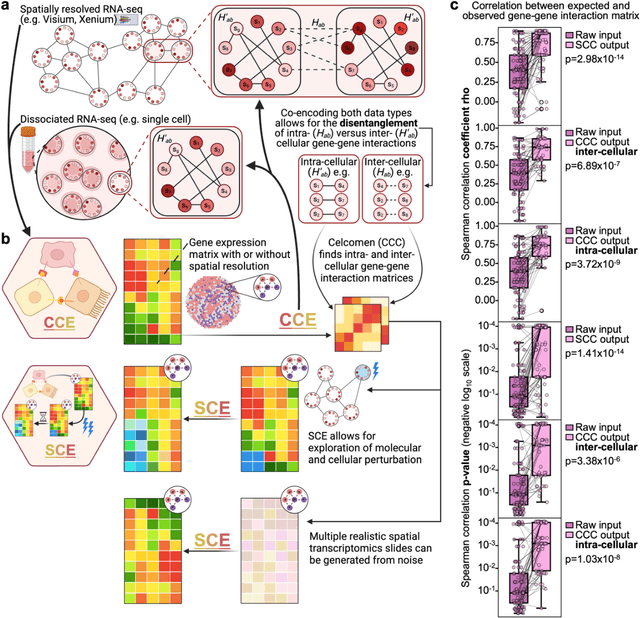
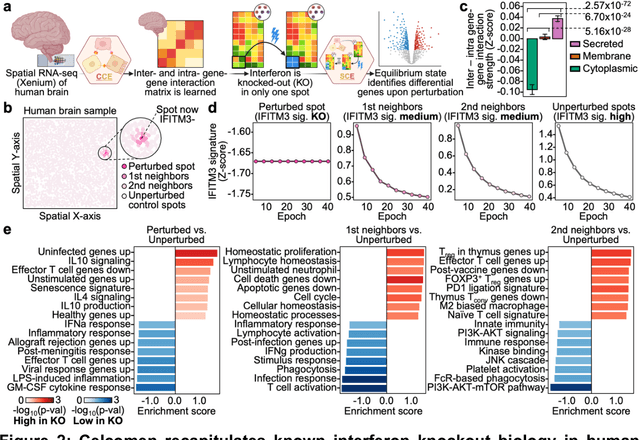
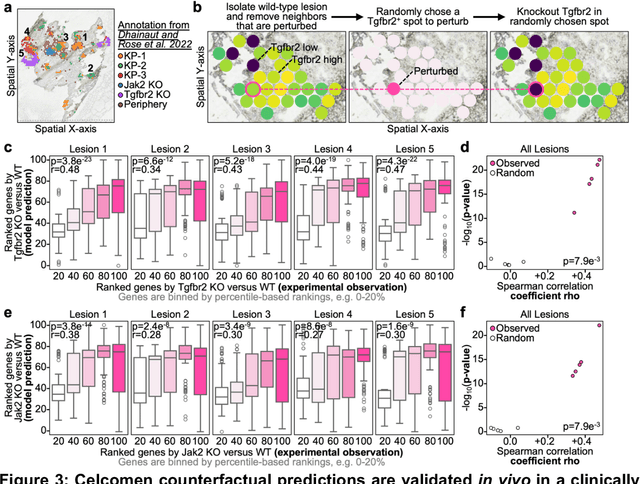
Abstract:Celcomen leverages a mathematical causality framework to disentangle intra- and inter- cellular gene regulation programs in spatial transcriptomics and single-cell data through a generative graph neural network. It can learn gene-gene interactions, as well as generate post-perturbation counterfactual spatial transcriptomics, thereby offering access to experimentally inaccessible samples. We validated its disentanglement, identifiability, and counterfactual prediction capabilities through simulations and in clinically relevant human glioblastoma, human fetal spleen, and mouse lung cancer samples. Celcomen provides the means to model disease and therapy induced changes allowing for new insights into single-cell spatially resolved tissue responses relevant to human health.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge