Christopher Irwin
HealthBranches: Synthesizing Clinically-Grounded Question Answering Datasets via Decision Pathways
Aug 10, 2025Abstract:HealthBranches is a novel benchmark dataset for medical Question-Answering (Q&A), specifically designed to evaluate complex reasoning in Large Language Models (LLMs). This dataset is generated through a semi-automated pipeline that transforms explicit decision pathways from medical source into realistic patient cases with associated questions and answers. Covering 4,063 case studies across 17 healthcare topics, each data point is based on clinically validated reasoning chains. HealthBranches supports both open-ended and multiple-choice question formats and uniquely includes the full reasoning path for each Q&A. Its structured design enables robust evaluation of LLMs' multi-step inference capabilities, including their performance in structured Retrieval-Augmented Generation (RAG) contexts. HealthBranches establishes a foundation for the development of more trustworthy, interpretable, and clinically reliable LLMs in high-stakes domains while also serving as a valuable resource for educational purposes.
Entropy-Lens: The Information Signature of Transformer Computations
Feb 23, 2025Abstract:Transformer models have revolutionized fields from natural language processing to computer vision, yet their internal computational dynamics remain poorly understood raising concerns about predictability and robustness. In this work, we introduce Entropy-Lens, a scalable, model-agnostic framework that leverages information theory to interpret frozen, off-the-shelf large-scale transformers. By quantifying the evolution of Shannon entropy within intermediate residual streams, our approach extracts computational signatures that distinguish model families, categorize task-specific prompts, and correlate with output accuracy. We further demonstrate the generality of our method by extending the analysis to vision transformers. Our results suggest that entropy-based metrics can serve as a principled tool for unveiling the inner workings of modern transformer architectures.
Beyond Labels: A Self-Supervised Framework with Masked Autoencoders and Random Cropping for Breast Cancer Subtype Classification
Oct 15, 2024


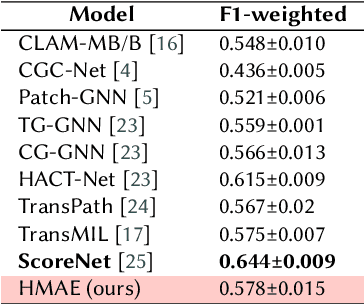
Abstract:This work contributes to breast cancer sub-type classification using histopathological images. We utilize masked autoencoders (MAEs) to learn a self-supervised embedding tailored for computer vision tasks in this domain. This embedding captures informative representations of histopathological data, facilitating feature learning without extensive labeled datasets. During pre-training, we investigate employing a random crop technique to generate a large dataset from WSIs automatically. Additionally, we assess the performance of linear probes for multi-class classification tasks of cancer sub-types using the representations learnt by the MAE. Our approach aims to achieve strong performance on downstream tasks by leveraging the complementary strengths of ViTs and autoencoders. We evaluate our model's performance on the BRACS dataset and compare it with existing benchmarks.
Graph Neural Networks for Gut Microbiome Metaomic data: A preliminary work
Jun 28, 2024

Abstract:The gut microbiome, crucial for human health, presents challenges in analyzing its complex metaomic data due to high dimensionality and sparsity. Traditional methods struggle to capture its intricate relationships. We investigate graph neural networks (GNNs) for this task, aiming to derive meaningful representations of individual gut microbiomes. Unlike methods relying solely on taxa abundance, we directly leverage phylogenetic relationships, in order to obtain a generalized encoder for taxa networks. The representation learnt from the encoder are then used to train a model for phenotype prediction such as Inflammatory Bowel Disease (IBD).
Structural Positional Encoding for knowledge integration in transformer-based medical process monitoring
Mar 13, 2024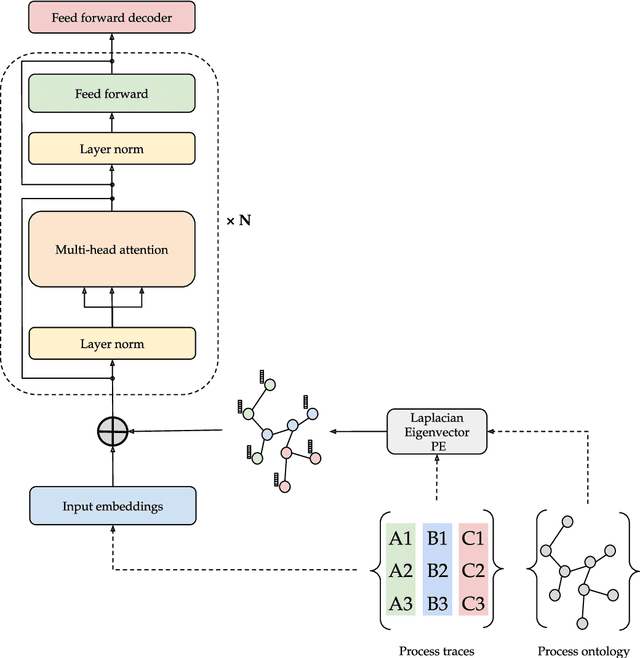



Abstract:Predictive process monitoring is a process mining task aimed at forecasting information about a running process trace, such as the most correct next activity to be executed. In medical domains, predictive process monitoring can provide valuable decision support in atypical and nontrivial situations. Decision support and quality assessment in medicine cannot ignore domain knowledge, in order to be grounded on all the available information (which is not limited to data) and to be really acceptable by end users. In this paper, we propose a predictive process monitoring approach relying on the use of a {\em transformer}, a deep learning architecture based on the attention mechanism. A major contribution of our work lies in the incorporation of ontological domain-specific knowledge, carried out through a graph positional encoding technique. The paper presents and discusses the encouraging experimental result we are collecting in the domain of stroke management.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge