Christopher Crane
Progressively refined deep joint registration segmentation (ProRSeg) of gastrointestinal organs at risk: Application to MRI and cone-beam CT
Oct 25, 2022



Abstract:Method: ProRSeg was trained using 5-fold cross-validation with 110 T2-weighted MRI acquired at 5 treatment fractions from 10 different patients, taking care that same patient scans were not placed in training and testing folds. Segmentation accuracy was measured using Dice similarity coefficient (DSC) and Hausdorff distance at 95th percentile (HD95). Registration consistency was measured using coefficient of variation (CV) in displacement of OARs. Ablation tests and accuracy comparisons against multiple methods were done. Finally, applicability of ProRSeg to segment cone-beam CT (CBCT) scans was evaluated on 80 scans using 5-fold cross-validation. Results: ProRSeg processed 3D volumes (128 $\times$ 192 $\times$ 128) in 3 secs on a NVIDIA Tesla V100 GPU. It's segmentations were significantly more accurate ($p<0.001$) than compared methods, achieving a DSC of 0.94 $\pm$0.02 for liver, 0.88$\pm$0.04 for large bowel, 0.78$\pm$0.03 for small bowel and 0.82$\pm$0.04 for stomach-duodenum from MRI. ProRSeg achieved a DSC of 0.72$\pm$0.01 for small bowel and 0.76$\pm$0.03 for stomach-duodenum from CBCT. ProRSeg registrations resulted in the lowest CV in displacement (stomach-duodenum $CV_{x}$: 0.75\%, $CV_{y}$: 0.73\%, and $CV_{z}$: 0.81\%; small bowel $CV_{x}$: 0.80\%, $CV_{y}$: 0.80\%, and $CV_{z}$: 0.68\%; large bowel $CV_{x}$: 0.71\%, $CV_{y}$ : 0.81\%, and $CV_{z}$: 0.75\%). ProRSeg based dose accumulation accounting for intra-fraction (pre-treatment to post-treatment MRI scan) and inter-fraction motion showed that the organ dose constraints were violated in 4 patients for stomach-duodenum and for 3 patients for small bowel. Study limitations include lack of independent testing and ground truth phantom datasets to measure dose accumulation accuracy.
Self-supervised 3D anatomy segmentation using self-distilled masked image transformer (SMIT)
May 20, 2022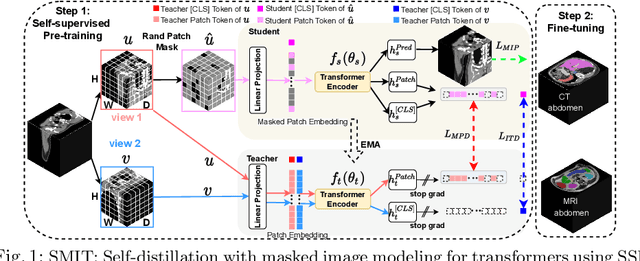
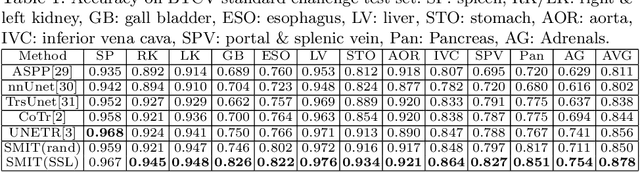
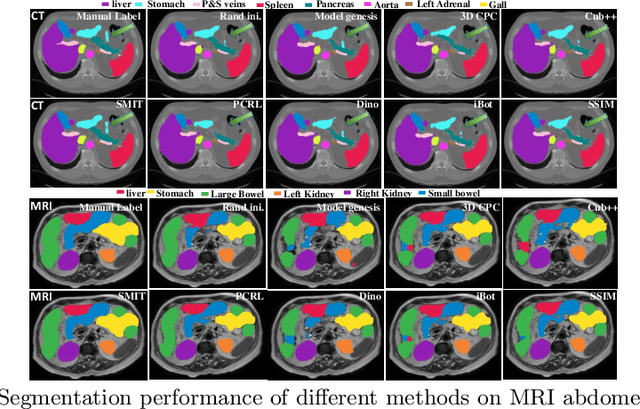
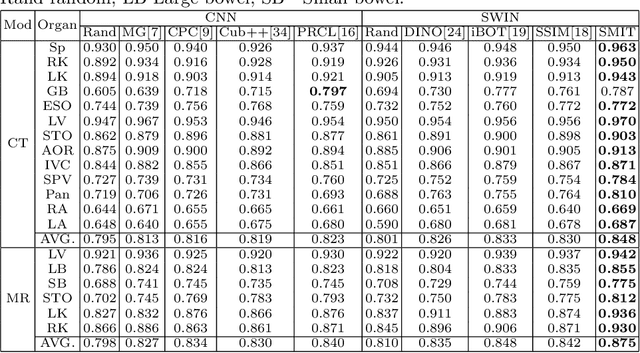
Abstract:Vision transformers, with their ability to more efficiently model long-range context, have demonstrated impressive accuracy gains in several computer vision and medical image analysis tasks including segmentation. However, such methods need large labeled datasets for training, which is hard to obtain for medical image analysis. Self-supervised learning (SSL) has demonstrated success in medical image segmentation using convolutional networks. In this work, we developed a \underline{s}elf-distillation learning with \underline{m}asked \underline{i}mage modeling method to perform SSL for vision \underline{t}ransformers (SMIT) applied to 3D multi-organ segmentation from CT and MRI. Our contribution is a dense pixel-wise regression within masked patches called masked image prediction, which we combined with masked patch token distillation as pretext task to pre-train vision transformers. We show our approach is more accurate and requires fewer fine tuning datasets than other pretext tasks. Unlike prior medical image methods, which typically used image sets arising from disease sites and imaging modalities corresponding to the target tasks, we used 3,643 CT scans (602,708 images) arising from head and neck, lung, and kidney cancers as well as COVID-19 for pre-training and applied it to abdominal organs segmentation from MRI pancreatic cancer patients as well as publicly available 13 different abdominal organs segmentation from CT. Our method showed clear accuracy improvement (average DSC of 0.875 from MRI and 0.878 from CT) with reduced requirement for fine-tuning datasets over commonly used pretext tasks. Extensive comparisons against multiple current SSL methods were done. Code will be made available upon acceptance for publication.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge