Christian John Hurry
Out-of-distribution evaluations of channel agnostic masked autoencoders in fluorescence microscopy
Mar 24, 2025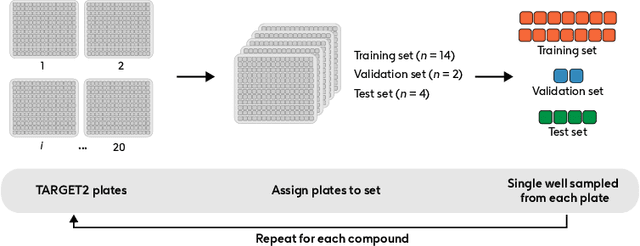
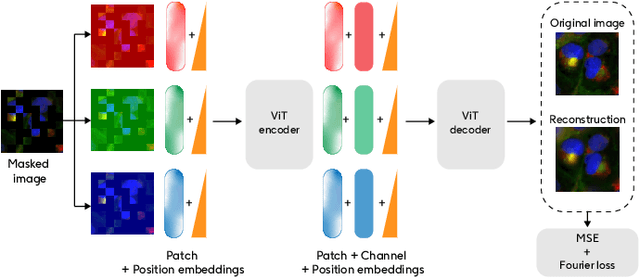
Abstract:Developing computer vision for high-content screening is challenging due to various sources of distribution-shift caused by changes in experimental conditions, perturbagens, and fluorescent markers. The impact of different sources of distribution-shift are confounded in typical evaluations of models based on transfer learning, which limits interpretations of how changes to model design and training affect generalisation. We propose an evaluation scheme that isolates sources of distribution-shift using the JUMP-CP dataset, allowing researchers to evaluate generalisation with respect to specific sources of distribution-shift. We then present a channel-agnostic masked autoencoder $\mathbf{Campfire}$ which, via a shared decoder for all channels, scales effectively to datasets containing many different fluorescent markers, and show that it generalises to out-of-distribution experimental batches, perturbagens, and fluorescent markers, and also demonstrates successful transfer learning from one cell type to another.
Self-supervised learning of multi-omics embeddings in the low-label, high-data regime
Nov 16, 2023



Abstract:Contrastive, self-supervised learning (SSL) is used to train a model that predicts cancer type from miRNA, mRNA or RPPA expression data. This model, a pretrained FT-Transformer, is shown to outperform XGBoost and CatBoost, standard benchmarks for tabular data, when labelled samples are scarce but the number of unlabelled samples is high. This is despite the fact that the datasets we use have $\mathcal{O}(10^{1})$ classes and $\mathcal{O}(10^{2})-\mathcal{O}(10^{4})$ features. After demonstrating the efficacy of our chosen method of self-supervised pretraining, we investigate SSL for multi-modal models. A late-fusion model is proposed, where each omics is passed through its own sub-network, the outputs of which are averaged and passed to the pretraining or downstream objective function. Multi-modal pretraining is shown to improve predictions from a single omics, and we argue that this is useful for datasets with many unlabelled multi-modal samples, but few labelled unimodal samples. Additionally, we show that pretraining each omics-specific module individually is highly effective. This enables the application of the proposed model in a variety of contexts where a large amount of unlabelled data is available from each omics, but only a few labelled samples.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge