Birgit Ertl-Wagner
Improving 3D convolutional neural network comprehensibility via interactive visualization of relevance maps: Evaluation in Alzheimer's disease
Dec 18, 2020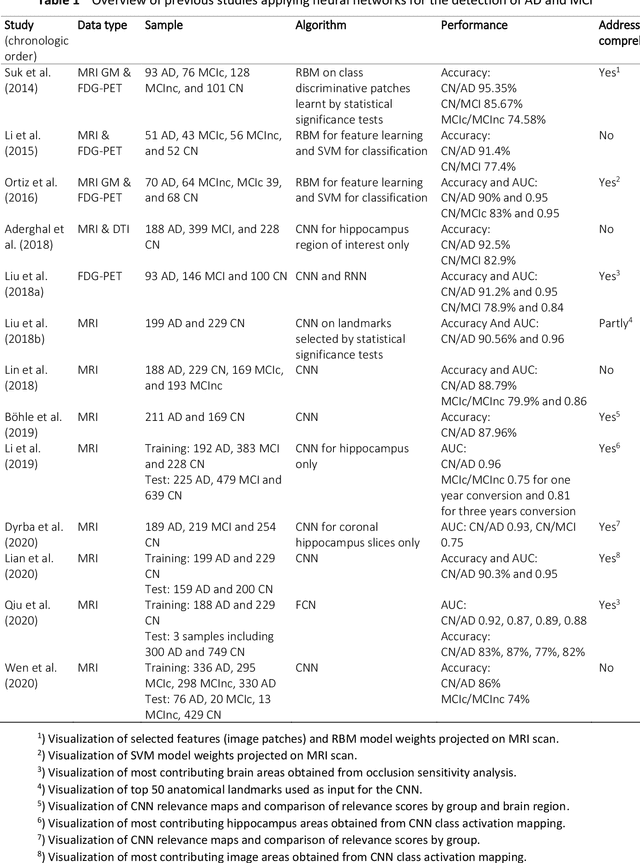
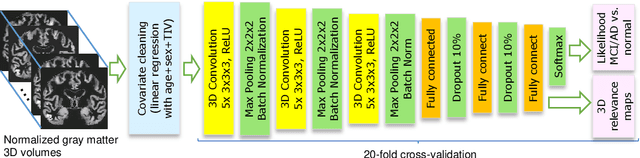
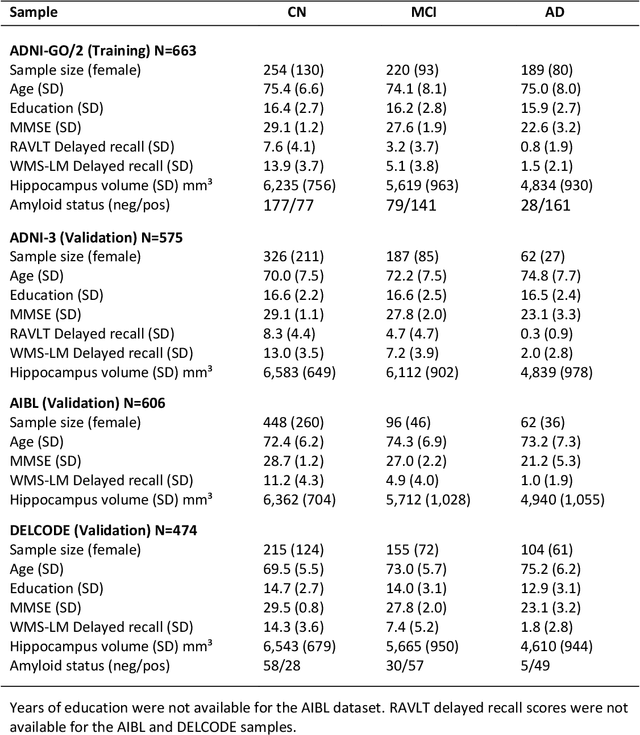
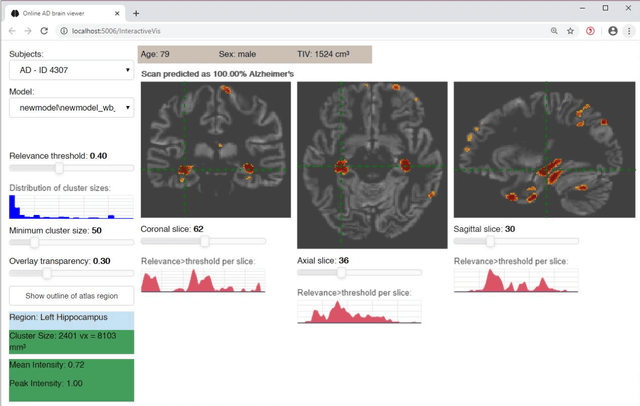
Abstract:Although convolutional neural networks (CNN) achieve high diagnostic accuracy for detecting Alzheimer's disease (AD) dementia based on magnetic resonance imaging (MRI) scans, they are not yet applied in clinical routine. One important reason for this is a lack of model comprehensibility. Recently developed visualization methods for deriving CNN relevance maps may help to fill this gap. We investigated whether models with higher accuracy also rely more on discriminative brain regions predefined by prior knowledge. We trained a CNN for the detection of AD in N=663 T1-weighted MRI scans of patients with dementia and amnestic mild cognitive impairment (MCI) and verified the accuracy of the models via cross-validation and in three independent samples including N=1655 cases. We evaluated the association of relevance scores and hippocampus volume to validate the clinical utility of this approach. To improve model comprehensibility, we implemented an interactive visualization of 3D CNN relevance maps. Across three independent datasets, group separation showed high accuracy for AD dementia vs. controls (AUC$\geq$0.92) and moderate accuracy for MCI vs. controls (AUC$\approx$0.75). Relevance maps indicated that hippocampal atrophy was considered as the most informative factor for AD detection, with additional contributions from atrophy in other cortical and subcortical regions. Relevance scores within the hippocampus were highly correlated with hippocampal volumes (Pearson's r$\approx$-0.81). The relevance maps highlighted atrophy in regions that we had hypothesized a priori. This strengthens the comprehensibility of the CNN models, which were trained in a purely data-driven manner based on the scans and diagnosis labels. The high hippocampus relevance scores and high performance achieved in independent samples support the validity of the CNN models in the detection of AD-related MRI abnormalities.
Hough-CNN: Deep Learning for Segmentation of Deep Brain Regions in MRI and Ultrasound
Jan 31, 2016
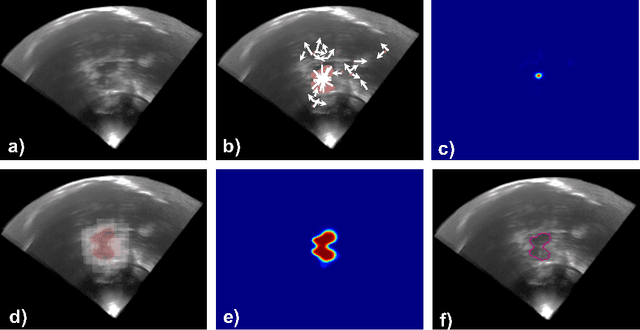

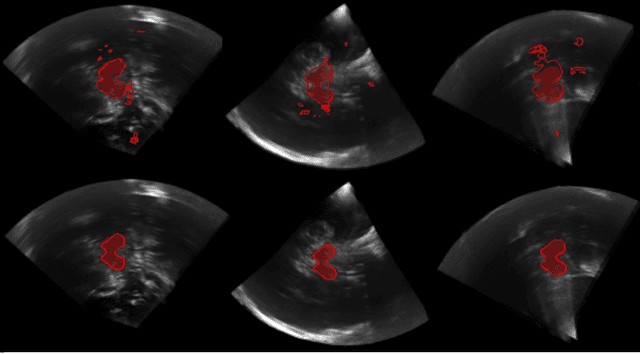
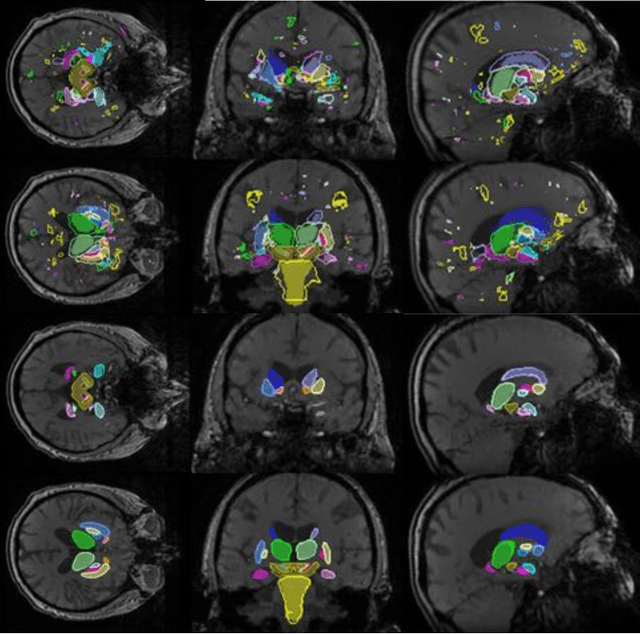
Abstract:In this work we propose a novel approach to perform segmentation by leveraging the abstraction capabilities of convolutional neural networks (CNNs). Our method is based on Hough voting, a strategy that allows for fully automatic localisation and segmentation of the anatomies of interest. This approach does not only use the CNN classification outcomes, but it also implements voting by exploiting the features produced by the deepest portion of the network. We show that this learning-based segmentation method is robust, multi-region, flexible and can be easily adapted to different modalities. In the attempt to show the capabilities and the behaviour of CNNs when they are applied to medical image analysis, we perform a systematic study of the performances of six different network architectures, conceived according to state-of-the-art criteria, in various situations. We evaluate the impact of both different amount of training data and different data dimensionality (2D, 2.5D and 3D) on the final results. We show results on both MRI and transcranial US volumes depicting respectively 26 regions of the basal ganglia and the midbrain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge