Askell Love
Automated brainstem parcellation using multi-atlas segmentation and deep neural network
Feb 05, 2021
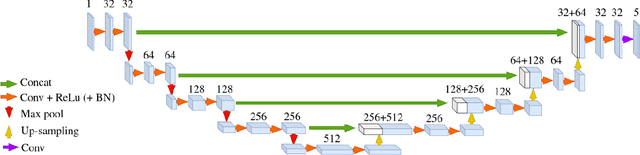
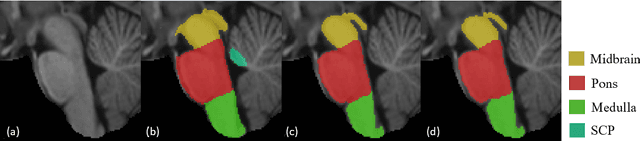
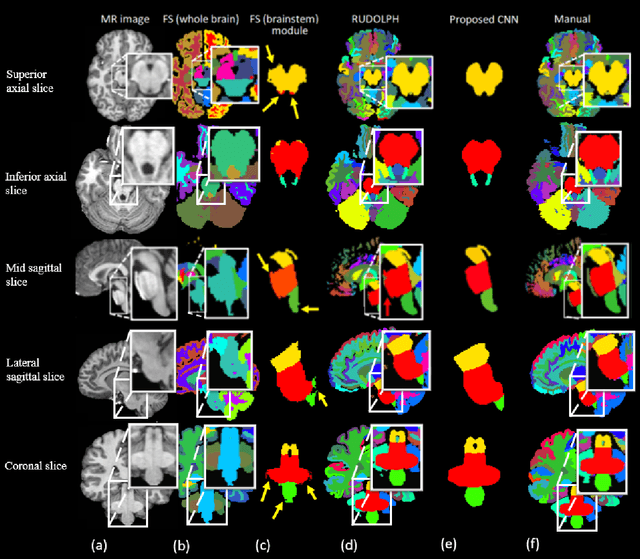
Abstract:About 5-8% of individuals over the age of 60 have dementia. With our ever-aging population this number is likely to increase, making dementia one of the most important threats to public health in the 21st century. Given the phenotypic overlap of individual dementias the diagnosis of dementia is a major clinical challenge, even with current gold standard diagnostic approaches. However, it has been shown that certain dementias show specific structural characteristics in the brain. Progressive supranuclear palsy (PSP) and multiple system atrophy (MSA) are prototypical examples of this phenomenon, as they often present with characteristic brainstem atrophy. More detailed characterization of brain atrophy due to individual diseases is urgently required to select biomarkers and therapeutic targets that are meaningful to each disease. Here we present a joint multi-atlas-segmentation and deep-learning-based segmentation method for fast and robust parcellation of the brainstem into its four sub-structures, i.e., the midbrain, pons, medulla, and superior cerebellar peduncles (SCP), that in turn can provide detailed volumetric information on the brainstem sub-structures affected in PSP and MSA. The method may also benefit other neurodegenerative diseases, such as Parkinson's disease; a condition which is often considered in the differential diagnosis of PSP and MSA. Comparison with state-of-the-art labeling techniques evaluated on ground truth manual segmentations demonstrate that our method is significantly faster than prior methods as well as showing improvement in labeling the brainstem indicating that this strategy may be a viable option to provide a better characterization of the brainstem atrophy seen in PSP and MSA.
Unsupervised brain lesion segmentation from MRI using a convolutional autoencoder
Nov 23, 2018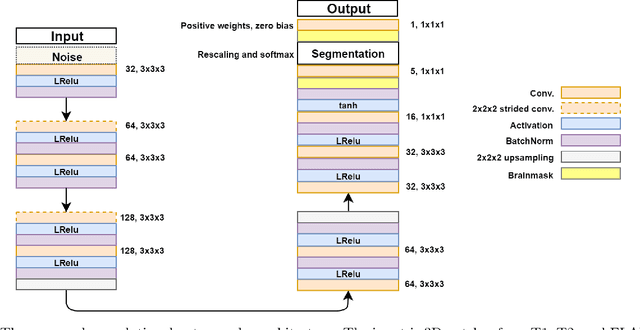
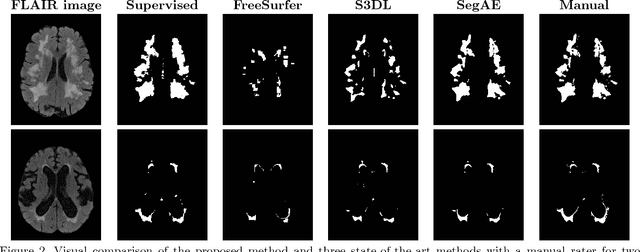

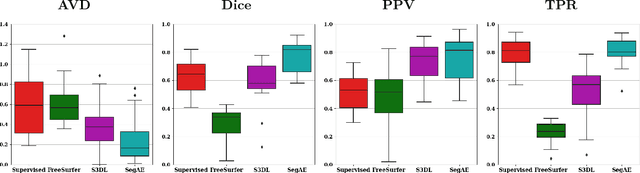
Abstract:Lesions that appear hyperintense in both Fluid Attenuated Inversion Recovery (FLAIR) and T2-weighted magnetic resonance images (MRIs) of the human brain are common in the brains of the elderly population and may be caused by ischemia or demyelination. Lesions are biomarkers for various neurodegenerative diseases, making accurate quantification of them important for both disease diagnosis and progression. Automatic lesion detection using supervised learning requires manually annotated images, which can often be impractical to acquire. Unsupervised lesion detection, on the other hand, does not require any manual delineation; however, these methods can be challenging to construct due to the variability in lesion load, placement of lesions, and voxel intensities. Here we present a novel approach to address this problem using a convolutional autoencoder, which learns to segment brain lesions as well as the white matter, gray matter, and cerebrospinal fluid by reconstructing FLAIR images as conical combinations of softmax layer outputs generated from the corresponding T1, T2, and FLAIR images. Some of the advantages of this model are that it accurately learns to segment lesions regardless of lesion load, and it can be used to quickly and robustly segment new images that were not in the training set. Comparisons with state-of-the-art segmentation methods evaluated on ground truth manual labels indicate that the proposed method works well for generating accurate lesion segmentations without the need for manual annotations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge