Ana Iglesias-Molina
EBOCA: Evidences for BiOmedical Concepts Association Ontology
Aug 01, 2022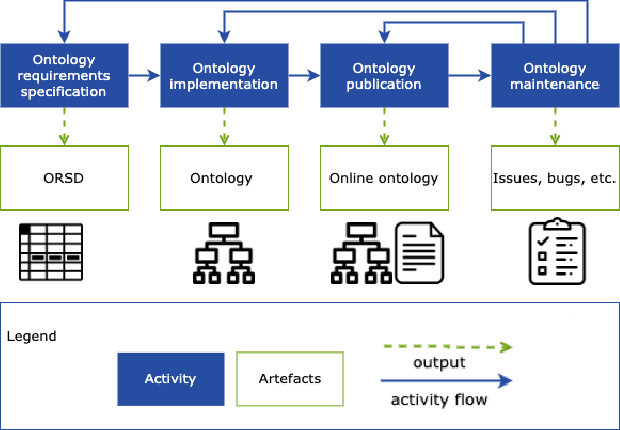
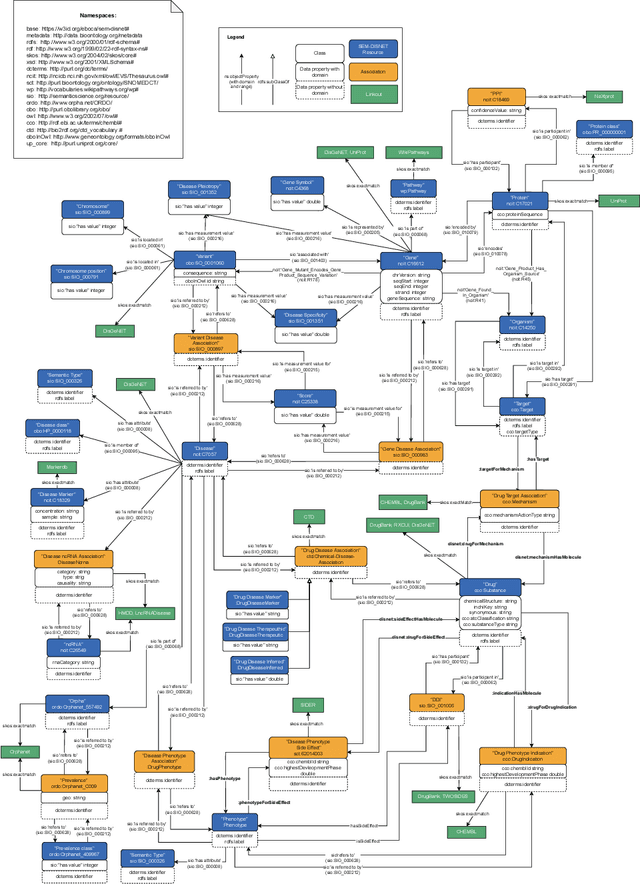
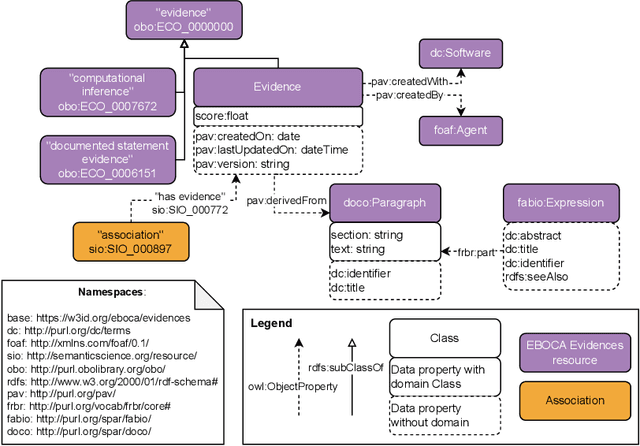
Abstract:There is a large number of online documents data sources available nowadays. The lack of structure and the differences between formats are the main difficulties to automatically extract information from them, which also has a negative impact on its use and reuse. In the biomedical domain, the DISNET platform emerged to provide researchers with a resource to obtain information in the scope of human disease networks by means of large-scale heterogeneous sources. Specifically in this domain, it is critical to offer not only the information extracted from different sources, but also the evidence that supports it. This paper proposes EBOCA, an ontology that describes (i) biomedical domain concepts and associations between them, and (ii) evidences supporting these associations; with the objective of providing an schema to improve the publication and description of evidences and biomedical associations in this domain. The ontology has been successfully evaluated to ensure there are no errors, modelling pitfalls and that it meets the previously defined functional requirements. Test data coming from a subset of DISNET and automatic association extractions from texts has been transformed according to the proposed ontology to create a Knowledge Graph that can be used in real scenarios, and which has also been used for the evaluation of the presented ontology.
Drugs4Covid: Drug-driven Knowledge Exploitation based on Scientific Publications
Dec 03, 2020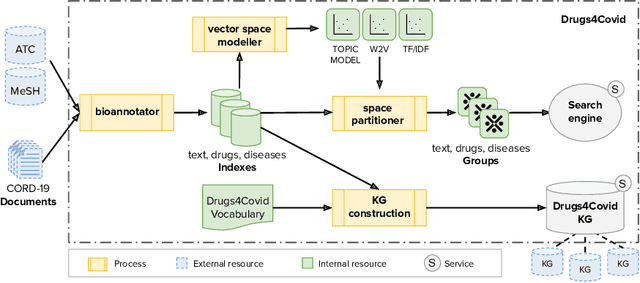
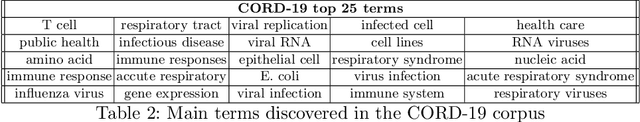
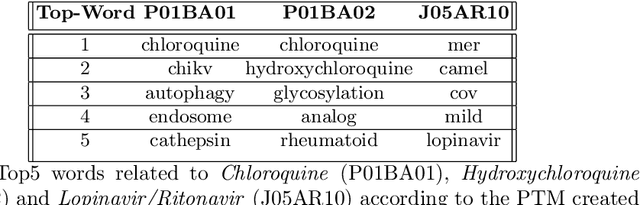


Abstract:In the absence of sufficient medication for COVID patients due to the increased demand, disused drugs have been employed or the doses of those available were modified by hospital pharmacists. Some evidences for the use of alternative drugs can be found in the existing scientific literature that could assist in such decisions. However, exploiting large corpus of documents in an efficient manner is not easy, since drugs may not appear explicitly related in the texts and could be mentioned under different brand names. Drugs4Covid combines word embedding techniques and semantic web technologies to enable a drug-oriented exploration of large medical literature. Drugs and diseases are identified according to the ATC classification and MeSH categories respectively. More than 60K articles and 2M paragraphs have been processed from the CORD-19 corpus with information of COVID-19, SARS, and other related coronaviruses. An open catalogue of drugs has been created and results are publicly available through a drug browser, a keyword-guided text explorer, and a knowledge graph.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge