Alison O'Neil
Controllable cardiac synthesis via disentangled anatomy arithmetic
Jul 04, 2021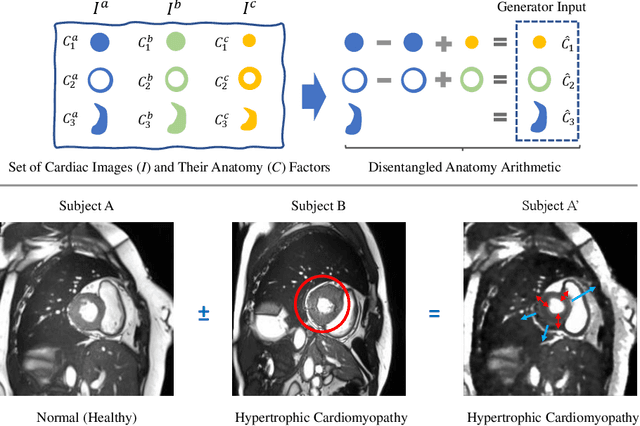
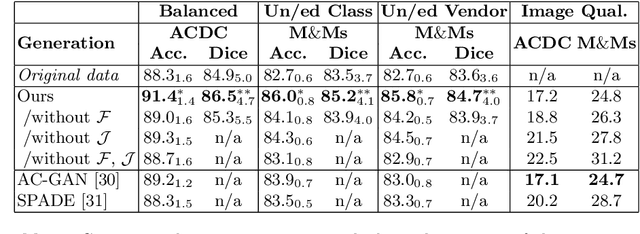
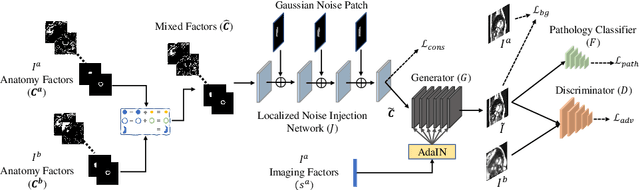

Abstract:Acquiring annotated data at scale with rare diseases or conditions remains a challenge. It would be extremely useful to have a method that controllably synthesizes images that can correct such underrepresentation. Assuming a proper latent representation, the idea of a "latent vector arithmetic" could offer the means of achieving such synthesis. A proper representation must encode the fidelity of the input data, preserve invariance and equivariance, and permit arithmetic operations. Motivated by the ability to disentangle images into spatial anatomy (tensor) factors and accompanying imaging (vector) representations, we propose a framework termed "disentangled anatomy arithmetic", in which a generative model learns to combine anatomical factors of different input images such that when they are re-entangled with the desired imaging modality (e.g. MRI), plausible new cardiac images are created with the target characteristics. To encourage a realistic combination of anatomy factors after the arithmetic step, we propose a localized noise injection network that precedes the generator. Our model is used to generate realistic images, pathology labels, and segmentation masks that are used to augment the existing datasets and subsequently improve post-hoc classification and segmentation tasks. Code is publicly available at https://github.com/vios-s/DAA-GAN.
Semi-supervised Meta-learning with Disentanglement for Domain-generalised Medical Image Segmentation
Jun 24, 2021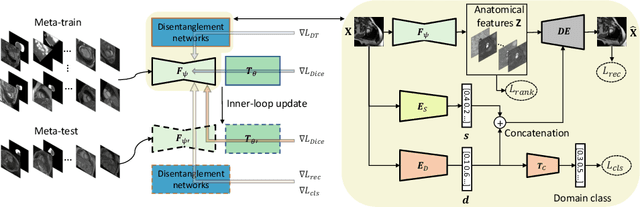
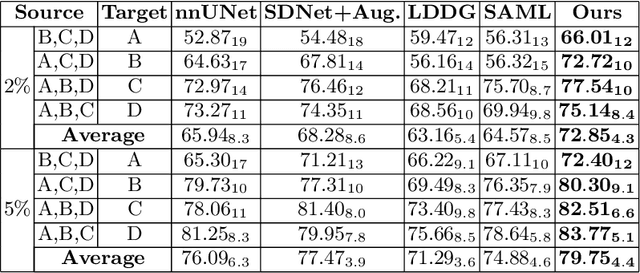

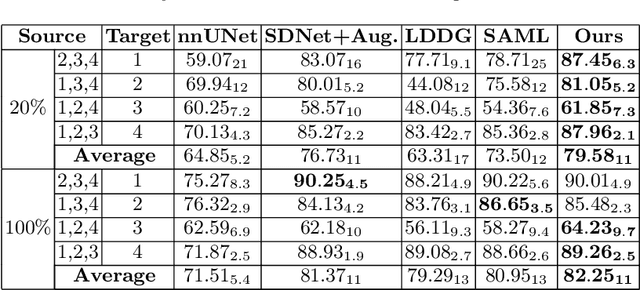
Abstract:Generalising deep models to new data from new centres (termed here domains) remains a challenge. This is largely attributed to shifts in data statistics (domain shifts) between source and unseen domains. Recently, gradient-based meta-learning approaches where the training data are split into meta-train and meta-test sets to simulate and handle the domain shifts during training have shown improved generalisation performance. However, the current fully supervised meta-learning approaches are not scalable for medical image segmentation, where large effort is required to create pixel-wise annotations. Meanwhile, in a low data regime, the simulated domain shifts may not approximate the true domain shifts well across source and unseen domains. To address this problem, we propose a novel semi-supervised meta-learning framework with disentanglement. We explicitly model the representations related to domain shifts. Disentangling the representations and combining them to reconstruct the input image allows unlabeled data to be used to better approximate the true domain shifts for meta-learning. Hence, the model can achieve better generalisation performance, especially when there is a limited amount of labeled data. Experiments show that the proposed method is robust on different segmentation tasks and achieves state-of-the-art generalisation performance on two public benchmarks.
Metrics for Exposing the Biases of Content-Style Disentanglement
Aug 31, 2020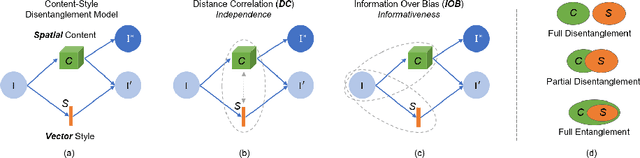
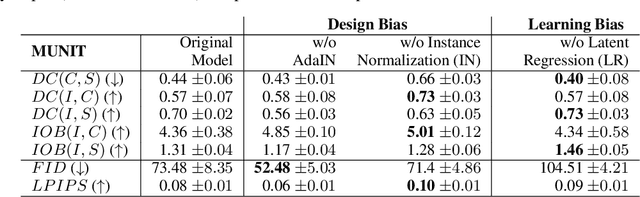
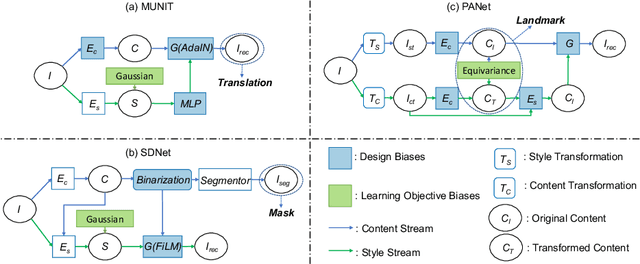

Abstract:Recent state-of-the-art semi- and un-supervised solutions for challenging computer vision tasks have used the idea of encoding image content into a spatial tensor and image appearance or "style" into a vector. These decomposed representations take advantage of equivariant properties of network design and improve performance in equivariant tasks, such as image-to-image translation. Most of these methods use the term "disentangled" for their representations and employ model design, learning objectives, and data biases to achieve good model performance. While considerable effort has been made to measure disentanglement in vector representations, currently, metrics that can characterize the degree of disentanglement between content (spatial) and style (vector) representations and the relation to task performance are lacking. In this paper, we propose metrics to measure how (un)correlated, biased, and informative the content and style representations are. In particular, we first identify key design choices and learning constraints on three popular models that employ content-style disentanglement and derive ablated versions. Then, we use our metrics to ascertain the role of each bias. Our experiments reveal a "sweet-spot" between disentanglement, task performance and latent space interpretability. The proposed metrics enable the design of better models and the selection of models that achieve the desired performance and disentanglement. Our metrics library is available at https://github.com/TsaftarisCollaboratory/CSDisentanglement_Metrics_Library.
Disentangled Representations for Domain-generalized Cardiac Segmentation
Aug 26, 2020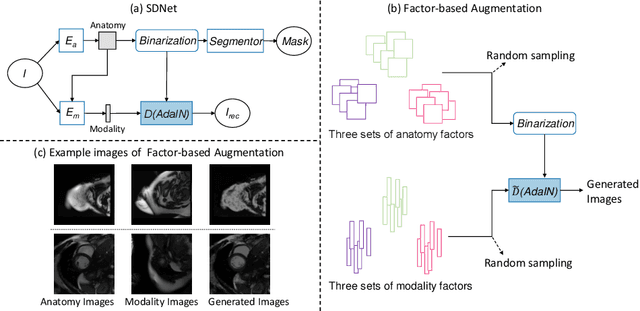

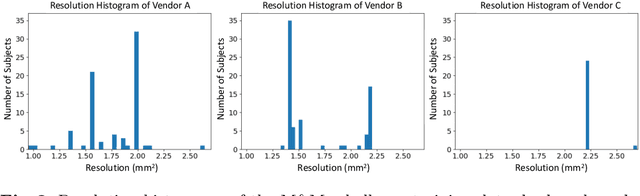
Abstract:Robust cardiac image segmentation is still an open challenge due to the inability of the existing methods to achieve satisfactory performance on unseen data of different domains. Since the acquisition and annotation of medical data are costly and time-consuming, recent work focuses on domain adaptation and generalization to bridge the gap between data from different populations and scanners. In this paper, we propose two data augmentation methods that focus on improving the domain adaptation and generalization abilities of state-to-the-art cardiac segmentation models. In particular, our "Resolution Augmentation" method generates more diverse data by rescaling images to different resolutions within a range spanning different scanner protocols. Subsequently, our "Factor-based Augmentation" method generates more diverse data by projecting the original samples onto disentangled latent spaces, and combining the learned anatomy and modality factors from different domains. Our extensive experiments demonstrate the importance of efficient adaptation between seen and unseen domains, as well as model generalization ability, to robust cardiac image segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge