Alan R. Lowe
Discovering interpretable models of scientific image data with deep learning
Feb 05, 2024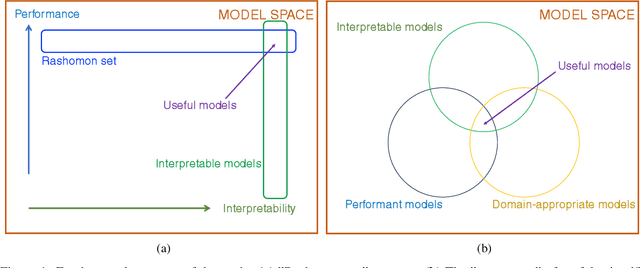
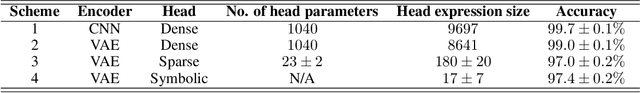
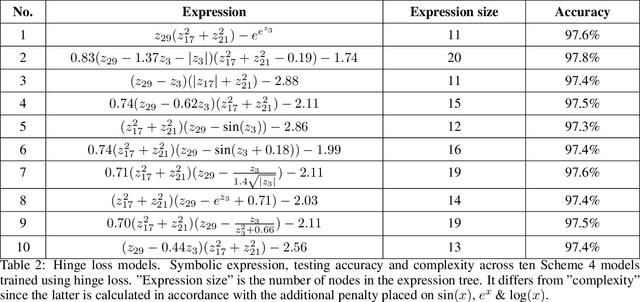



Abstract:How can we find interpretable, domain-appropriate models of natural phenomena given some complex, raw data such as images? Can we use such models to derive scientific insight from the data? In this paper, we propose some methods for achieving this. In particular, we implement disentangled representation learning, sparse deep neural network training and symbolic regression, and assess their usefulness in forming interpretable models of complex image data. We demonstrate their relevance to the field of bioimaging using a well-studied test problem of classifying cell states in microscopy data. We find that such methods can produce highly parsimonious models that achieve $\sim98\%$ of the accuracy of black-box benchmark models, with a tiny fraction of the complexity. We explore the utility of such interpretable models in producing scientific explanations of the underlying biological phenomenon.
Affinity-VAE for disentanglement, clustering and classification of objects in multidimensional image data
Sep 09, 2022

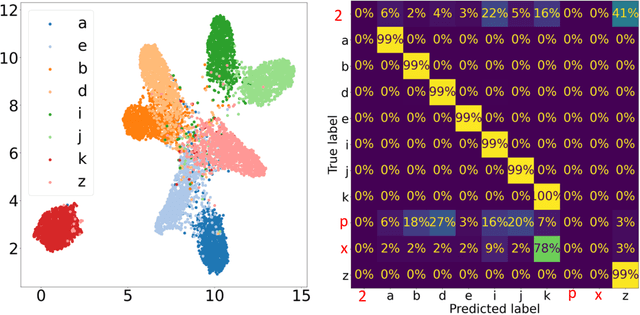

Abstract:In this work we present affinity-VAE: a framework for automatic clustering and classification of objects in multidimensional image data based on their similarity. The method expands on the concept of $\beta$-VAEs with an informed similarity-based loss component driven by an affinity matrix. The affinity-VAE is able to create rotationally-invariant, morphologically homogeneous clusters in the latent representation, with improved cluster separation compared with a standard $\beta$-VAE. We explore the extent of latent disentanglement and continuity of the latent spaces on both 2D and 3D image data, including simulated biological electron cryo-tomography (cryo-ET) volumes as an example of a scientific application.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge