Adriana V. Gregory
Developing a Machine Learning-Based Clinical Decision Support Tool for Uterine Tumor Imaging
Aug 20, 2023



Abstract:Uterine leiomyosarcoma (LMS) is a rare but aggressive malignancy. On imaging, it is difficult to differentiate LMS from, for example, degenerated leiomyoma (LM), a prevalent but benign condition. We curated a data set of 115 axial T2-weighted MRI images from 110 patients (mean [range] age=45 [17-81] years) with UTs that included five different tumor types. These data were randomly split stratifying on tumor volume into training (n=85) and test sets (n=30). An independent second reader (reader 2) provided manual segmentations for all test set images. To automate segmentation, we applied nnU-Net and explored the effect of training set size on performance by randomly generating subsets with 25, 45, 65 and 85 training set images. We evaluated the ability of radiomic features to distinguish between types of UT individually and when combined through feature selection and machine learning. Using the entire training set the mean [95% CI] fibroid DSC was measured as 0.87 [0.59-1.00] and the agreement between the two readers was 0.89 [0.77-1.0] on the test set. When classifying degenerated LM from LMS we achieve a test set F1-score of 0.80. Classifying UTs based on radiomic features we identify classifiers achieving F1-scores of 0.53 [0.45, 0.61] and 0.80 [0.80, 0.80] on the test set for the benign versus malignant, and degenerated LM versus LMS tasks. We show that it is possible to develop an automated method for 3D segmentation of the uterus and UT that is close to human-level performance with fewer than 150 annotated images. For distinguishing UT types, while we train models that merit further investigation with additional data, reliable automatic differentiation of UTs remains a challenge.
Role of Image Acquisition and Patient Phenotype Variations in Automatic Segmentation Model Generalization
Jul 26, 2023



Abstract:Purpose: This study evaluated the out-of-domain performance and generalization capabilities of automated medical image segmentation models, with a particular focus on adaptation to new image acquisitions and disease type. Materials: Datasets from both non-contrast and contrast-enhanced abdominal CT scans of healthy patients and those with polycystic kidney disease (PKD) were used. A total of 400 images (100 non-contrast controls, 100 contrast controls, 100 non-contrast PKD, 100 contrast PKD) were utilized for training/validation of models to segment kidneys, livers, and spleens, and the final models were then tested on 100 non-contrast CT images of patients affected by PKD. Performance was evaluated using Dice, Jaccard, TPR, and Precision. Results: Models trained on a diverse range of data showed no worse performance than models trained exclusively on in-domain data when tested on in-domain data. For instance, the Dice similarity of the model trained on 25% from each dataset was found to be non-inferior to the model trained purely on in-domain data. Conclusions: The results indicate that broader training examples significantly enhances model generalization and out-of-domain performance, thereby improving automated segmentation tools' applicability in clinical settings. The study's findings provide a roadmap for future research to adopt a data-centric approach in medical image AI model development.
AI in the Loop -- Functionalizing Fold Performance Disagreement to Monitor Automated Medical Image Segmentation Pipelines
May 15, 2023

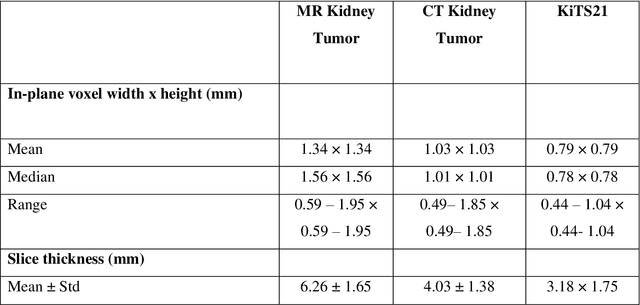

Abstract:Methods for automatically flag poor performing-predictions are essential for safely implementing machine learning workflows into clinical practice and for identifying difficult cases during model training. We present a readily adoptable method using sub-models trained on different dataset folds, where their disagreement serves as a surrogate for model confidence. Thresholds informed by human interobserver values were used to determine whether a final ensemble model prediction would require manual review. In two different datasets (abdominal CT and MR predicting kidney tumors), our framework effectively identified low performing automated segmentations. Flagging images with a minimum Interfold test Dice score below human interobserver variability maximized the number of flagged images while ensuring maximum ensemble test Dice. When our internally trained model was applied to an external publicly available dataset (KiTS21), flagged images included smaller tumors than those observed in our internally trained dataset, demonstrating the methods robustness to flagging poor performing out-of-distribution input data. Comparing interfold sub-model disagreement against human interobserver values is an efficient way to approximate a model's epistemic uncertainty - its lack of knowledge due to insufficient relevant training data - a key functionality for adopting these applications in clinical practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge