Utilizing Uncertainty Estimation in Deep Learning Segmentation of Fluorescence Microscopy Images with Missing Markers
Paper and Code
Jan 27, 2021
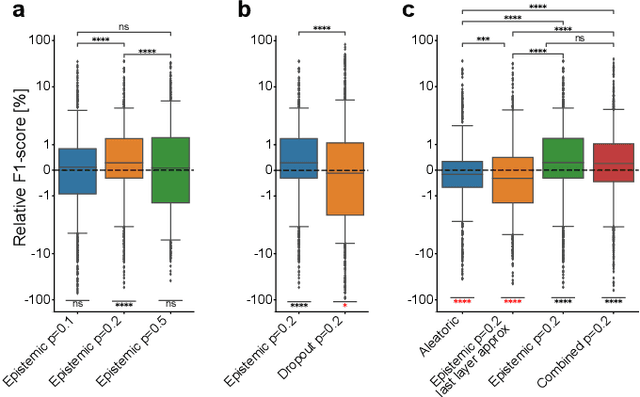
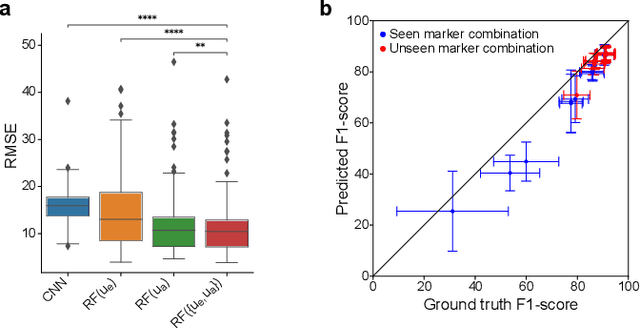
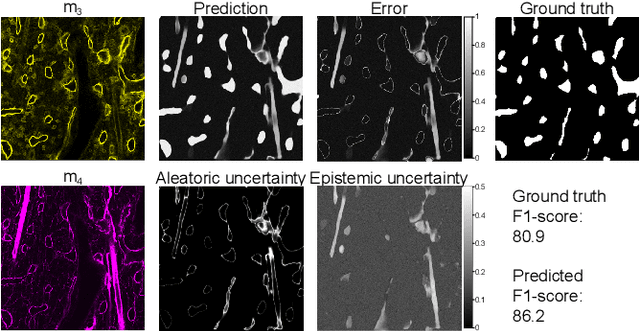
Fluorescence microscopy images contain several channels, each indicating a marker staining the sample. Since many different marker combinations are utilized in practice, it has been challenging to apply deep learning based segmentation models, which expect a predefined channel combination for all training samples as well as at inference for future application. Recent work circumvents this problem using a modality attention approach to be effective across any possible marker combination. However, for combinations that do not exist in a labeled training dataset, one cannot have any estimation of potential segmentation quality if that combination is encountered during inference. Without this, not only one lacks quality assurance but one also does not know where to put any additional imaging and labeling effort. We herein propose a method to estimate segmentation quality on unlabeled images by (i) estimating both aleatoric and epistemic uncertainties of convolutional neural networks for image segmentation, and (ii) training a Random Forest model for the interpretation of uncertainty features via regression to their corresponding segmentation metrics. Additionally, we demonstrate that including these uncertainty measures during training can provide an improvement on segmentation performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge