Unsupervised Reverse Domain Adaptation for Synthetic Medical Images via Adversarial Training
Paper and Code
Nov 29, 2017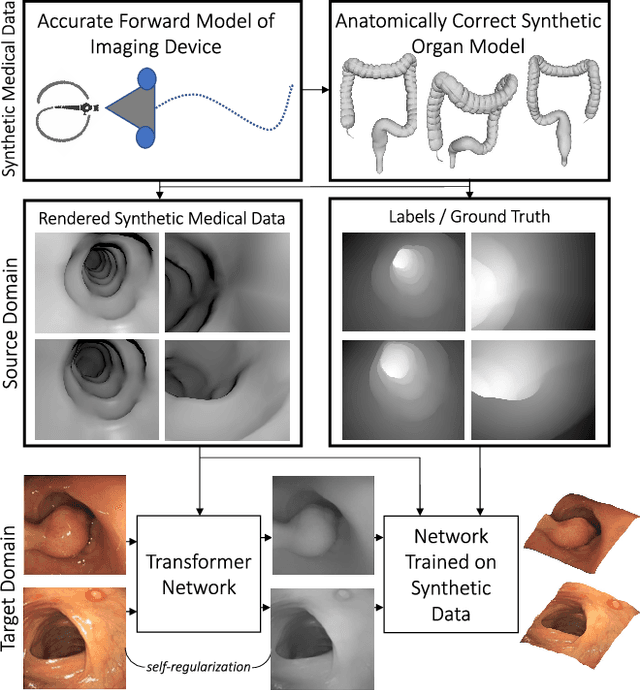



To realize the full potential of deep learning for medical imaging, large annotated datasets are required for training. Such datasets are difficult to acquire because labeled medical images are not usually available due to privacy issues, lack of experts available for annotation, underrepresentation of rare conditions and poor standardization. Lack of annotated data has been addressed in conventional vision applications using synthetic images refined via unsupervised adversarial training to look like real images. However, this approach is difficult to extend to general medical imaging because of the complex and diverse set of features found in real human tissues. We propose an alternative framework that uses a reverse flow, where adversarial training is used to make real medical images more like synthetic images, and hypothesize that clinically-relevant features can be preserved via self-regularization. These domain-adapted images can then be accurately interpreted by networks trained on large datasets of synthetic medical images. We test this approach for the notoriously difficult task of depth-estimation from endoscopy. We train a depth estimator on a large dataset of synthetic images generated using an accurate forward model of an endoscope and an anatomically-realistic colon. This network predicts significantly better depths when using synthetic-like domain-adapted images compared to the real images, confirming that the clinically-relevant features of depth are preserved.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge