TacoGFN: Target Conditioned GFlowNet for Structure-Based Drug Design
Paper and Code
Oct 05, 2023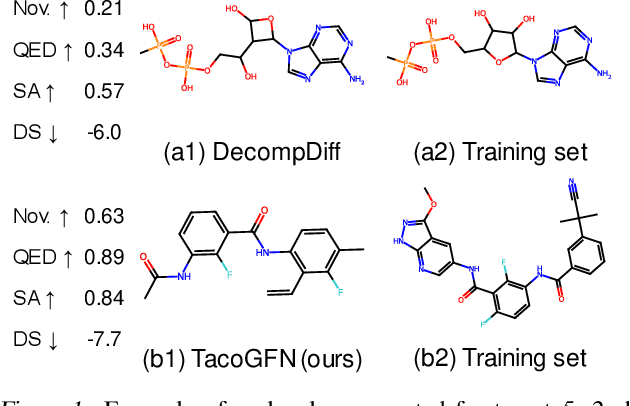



We seek to automate the generation of drug-like compounds conditioned to specific protein pocket targets. Most current methods approximate the protein-molecule distribution of a finite dataset and, therefore struggle to generate molecules with significant binding improvement over the training dataset. We instead frame the pocket-conditioned molecular generation task as an RL problem and develop TacoGFN, a target conditional Generative Flow Network model. Our method is explicitly encouraged to generate molecules with desired properties as opposed to fitting on a pre-existing data distribution. To this end, we develop transformer-based docking score prediction to speed up docking score computation and propose TacoGFN to explore molecule space efficiently. Furthermore, we incorporate several rounds of active learning where generated samples are queried using a docking oracle to improve the docking score prediction. This approach allows us to accurately explore as much of the molecule landscape as we can afford computationally. Empirically, molecules generated using TacoGFN and its variants significantly outperform all baseline methods across every property (Docking score, QED, SA, Lipinski), while being orders of magnitude faster.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge