Sparse evolutionary Deep Learning with over one million artificial neurons on commodity hardware
Paper and Code
Jan 26, 2019
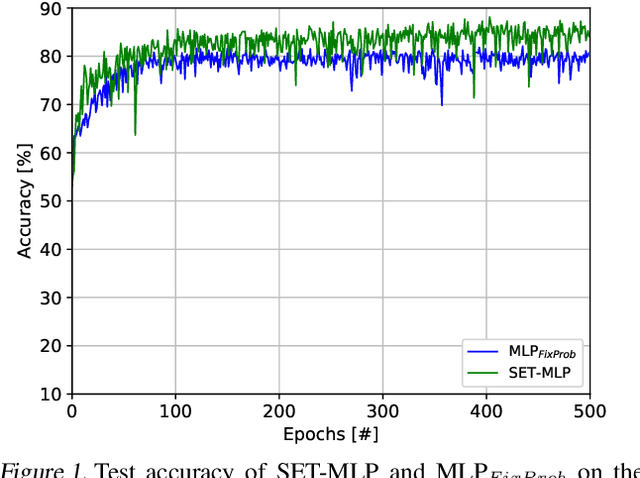
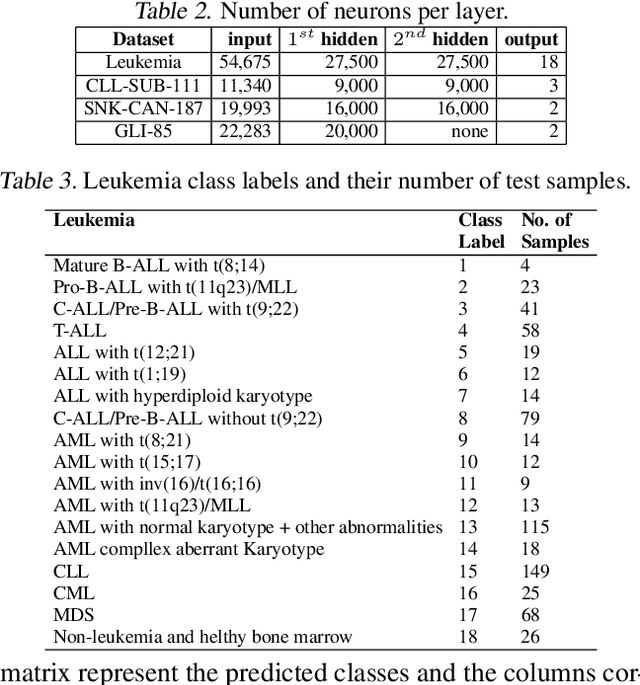
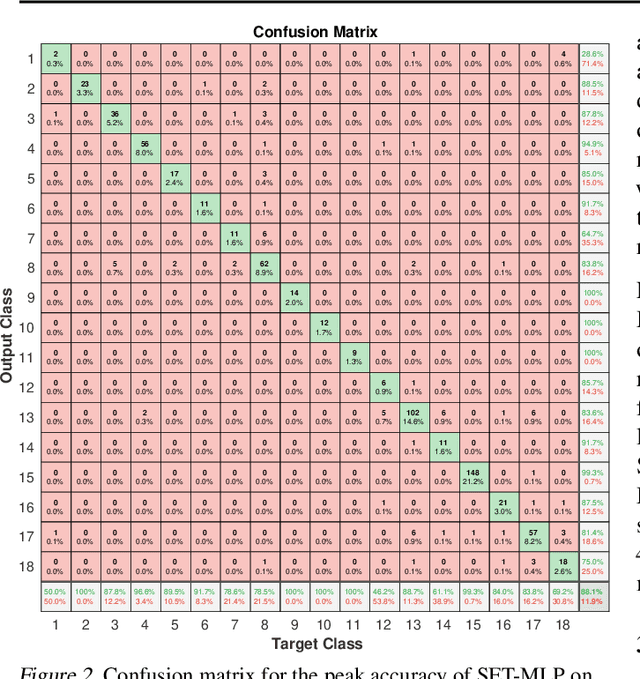
Microarray gene expression has widely attracted the eyes of the public as an efficient tool for cancer diagnosis and classification. However, the very-high dimensionality and the small number of samples make it difficult for traditional machine learning algorithms to address this problem due to the high amount of computations required and overfitting. So far, the existing approaches of processing microarray datasets are still far from satisfactory and they employ two phases, feature selection (or extraction) followed by a machine learning algorithm. In this paper, we show that MultiLayer Perceptrons (MLPs) with adaptive sparse connectivity can directly handle this problem without features selection. Tested on four datasets, our novel results demonstrate that deep learning methods can be applied directly also to high dimensional non-grid like data, while learning from a small amount of labeled examples with imbalanced classes and achieving better accuracy than the traditional two phases approach. Moreover, we have been able to create sparse MLP models with over one million neurons and to train them on a typical laptop without GPU. This is with two orders of magnitude more than the largest MLPs which can run currently on commodity hardware.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge