RGCVAE: Relational Graph Conditioned Variational Autoencoder for Molecule Design
Paper and Code
May 19, 2023
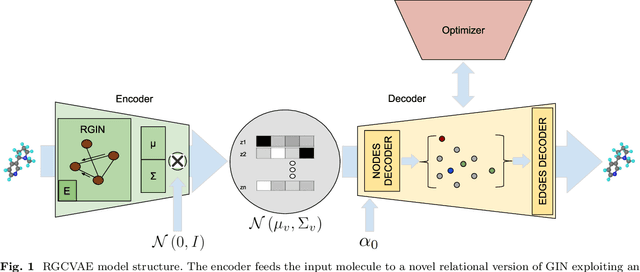
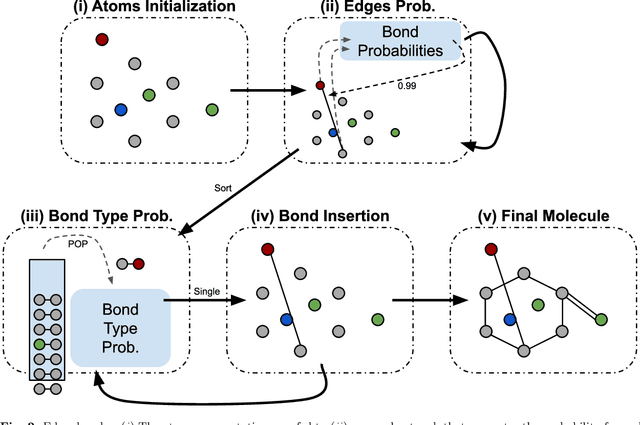
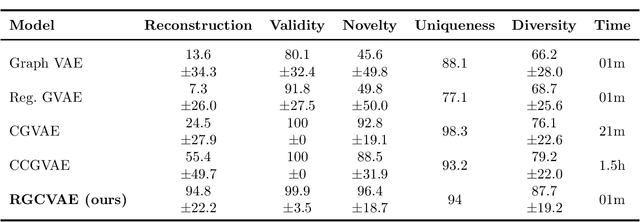
Identifying molecules that exhibit some pre-specified properties is a difficult problem to solve. In the last few years, deep generative models have been used for molecule generation. Deep Graph Variational Autoencoders are among the most powerful machine learning tools with which it is possible to address this problem. However, existing methods struggle in capturing the true data distribution and tend to be computationally expensive. In this work, we propose RGCVAE, an efficient and effective Graph Variational Autoencoder based on: (i) an encoding network exploiting a new powerful Relational Graph Isomorphism Network; (ii) a novel probabilistic decoding component. Compared to several state-of-the-art VAE methods on two widely adopted datasets, RGCVAE shows state-of-the-art molecule generation performance while being significantly faster to train.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge