Principled Hyperedge Prediction with Structural Spectral Features and Neural Networks
Paper and Code
Jun 13, 2021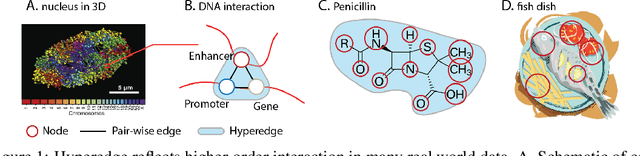

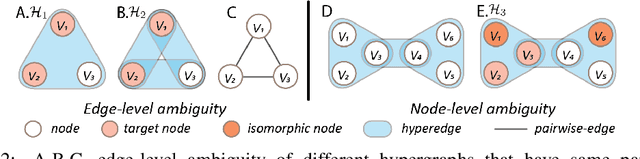

Hypergraph offers a framework to depict the multilateral relationships in real-world complex data. Predicting higher-order relationships, i.e hyperedge, becomes a fundamental problem for the full understanding of complicated interactions. The development of graph neural network (GNN) has greatly advanced the analysis of ordinary graphs with pair-wise relations. However, these methods could not be easily extended to the case of hypergraph. In this paper, we generalize the challenges of GNN in representing higher-order data in principle, which are edge- and node-level ambiguities. To overcome the challenges, we present SNALS that utilizes bipartite graph neural network with structural features to collectively tackle the two ambiguity issues. SNALS captures the joint interactions of a hyperedge by its local environment, which is retrieved by collecting the spectrum information of their connections. As a result, SNALS achieves nearly 30% performance increase compared with most recent GNN-based models. In addition, we applied SNALS to predict genetic higher-order interactions on 3D genome organization data. SNALS showed consistently high prediction accuracy across different chromosomes, and generated novel findings on 4-way gene interaction, which is further validated by existing literature.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge