Preserved Edge Convolutional Neural Network for Sensitivity Enhancement of Deuterium Metabolic Imaging (DMI)
Paper and Code
Sep 13, 2023
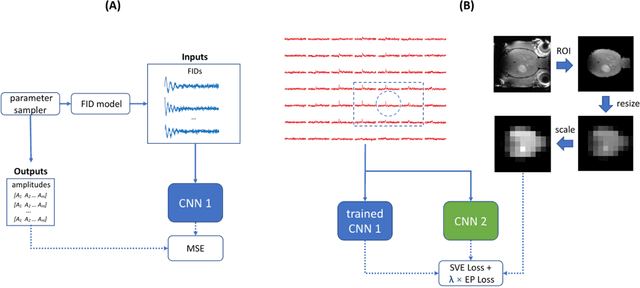
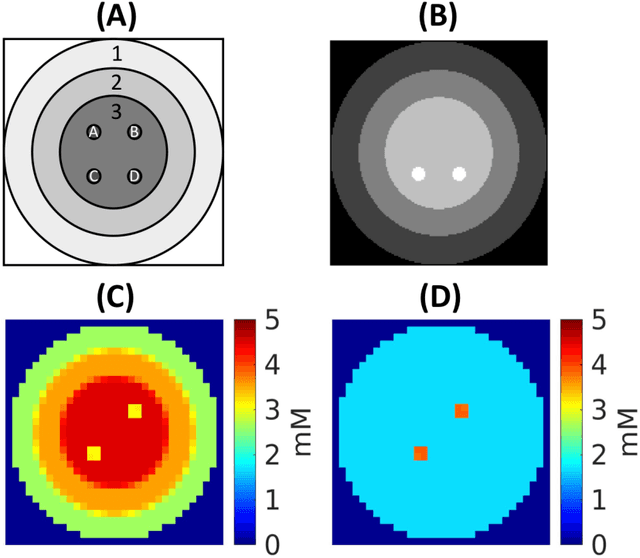
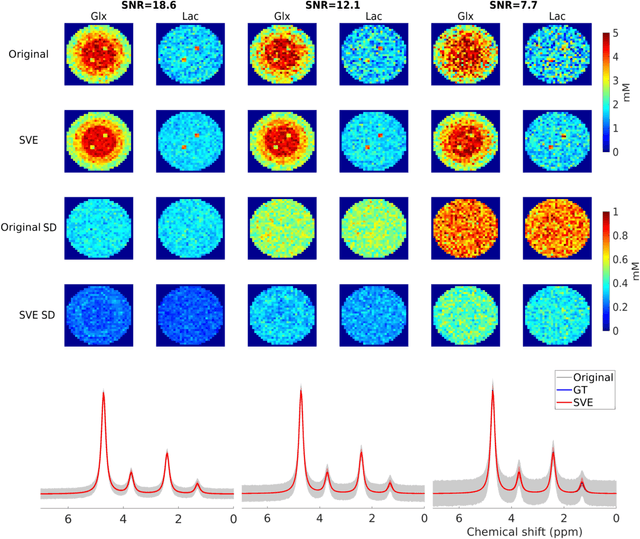
Purpose: Common to most MRSI techniques, the spatial resolution and the minimal scan duration of Deuterium Metabolic Imaging (DMI) are limited by the achievable SNR. This work presents a deep learning method for sensitivity enhancement of DMI. Methods: A convolutional neural network (CNN) was designed to estimate the 2H-labeled metabolite concentrations from low SNR and distorted DMI FIDs. The CNN was trained with synthetic data that represent a range of SNR levels typically encountered in vivo. The estimation precision was further improved by fine-tuning the CNN with MRI-based edge-preserving regularization for each DMI dataset. The proposed processing method, PReserved Edge ConvolutIonal neural network for Sensitivity Enhanced DMI (PRECISE-DMI), was applied to simulation studies and in vivo experiments to evaluate the anticipated improvements in SNR and investigate the potential for inaccuracies. Results: PRECISE-DMI visually improved the metabolic maps of low SNR datasets, and quantitatively provided higher precision than the standard Fourier reconstruction. Processing of DMI data acquired in rat brain tumor models resulted in more precise determination of 2H-labeled lactate and glutamate + glutamine levels, at increased spatial resolution (from >8 to 2 $\mu$L) or shortened scan time (from 32 to 4 min) compared to standard acquisitions. However, rigorous SD-bias analyses showed that overuse of the edge-preserving regularization can compromise the accuracy of the results. Conclusion: PRECISE-DMI allows a flexible trade-off between enhancing the sensitivity of DMI and minimizing the inaccuracies. With typical settings, the DMI sensitivity can be improved by 3-fold while retaining the capability to detect local signal variations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge