Mew: Multiplexed Immunofluorescence Image Analysis through an Efficient Multiplex Network
Paper and Code
Jul 25, 2024

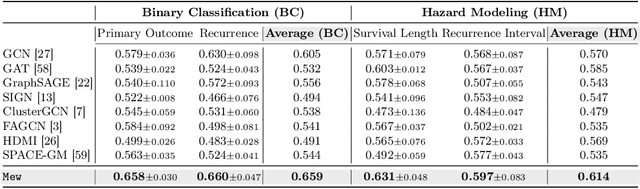

Recent advancements in graph-based approaches for multiplexed immunofluorescence (mIF) images have significantly propelled the field forward, offering deeper insights into patient-level phenotyping. However, current graph-based methodologies encounter two primary challenges: (1) Cellular Heterogeneity, where existing approaches fail to adequately address the inductive biases inherent in graphs, particularly the homophily characteristic observed in cellular connectivity and; (2) Scalability, where handling cellular graphs from high-dimensional images faces difficulties in managing a high number of cells. To overcome these limitations, we introduce Mew, a novel framework designed to efficiently process mIF images through the lens of multiplex network. Mew innovatively constructs a multiplex network comprising two distinct layers: a Voronoi network for geometric information and a Cell-type network for capturing cell-wise homogeneity. This framework equips a scalable and efficient Graph Neural Network (GNN), capable of processing the entire graph during training. Furthermore, Mew integrates an interpretable attention module that autonomously identifies relevant layers for image classification. Extensive experiments on a real-world patient dataset from various institutions highlight Mew's remarkable efficacy and efficiency, marking a significant advancement in mIF image analysis. The source code of Mew can be found here: \url{https://github.com/UNITES-Lab/Mew}
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge