METCC: METric learning for Confounder Control Making distance matter in high dimensional biological analysis
Paper and Code
Dec 07, 2018
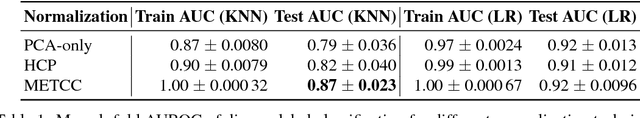
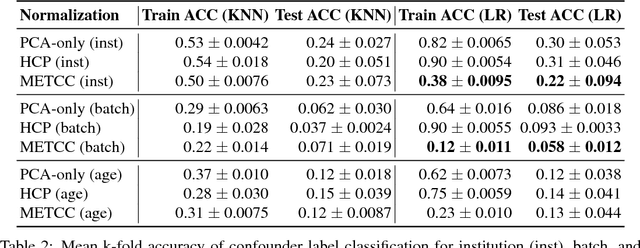
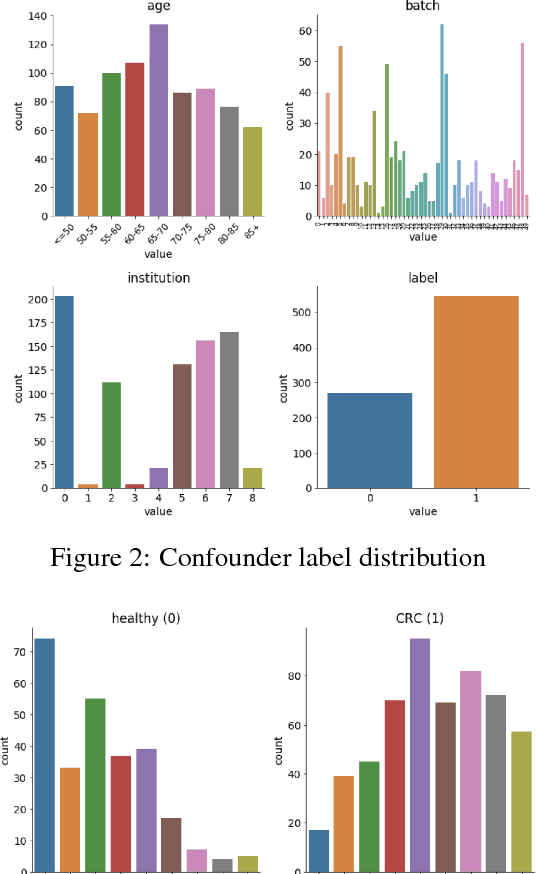
High-dimensional data acquired from biological experiments such as next generation sequencing are subject to a number of confounding effects. These effects include both technical effects, such as variation across batches from instrument noise or sample processing, or institution-specific differences in sample acquisition and physical handling, as well as biological effects arising from true but irrelevant differences in the biology of each sample, such as age biases in diseases. Prior work has used linear methods to adjust for such batch effects. Here, we apply contrastive metric learning by a non-linear triplet network to optimize the ability to distinguish biologically distinct sample classes in the presence of irrelevant technical and biological variation. Using whole-genome cell-free DNA data from 817 patients, we demonstrate that our approach, METric learning for Confounder Control (METCC), is able to match or exceed the classification performance achieved using a best-in-class linear method (HCP) or no normalization. Critically, results from METCC appear less confounded by irrelevant technical variables like institution and batch than those from other methods even without access to high quality metadata information required by many existing techniques; offering hope for improved generalization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge