Machine Learning-Based Analysis of Ebola Virus' Impact on Gene Expression in Nonhuman Primates
Paper and Code
Jan 22, 2024
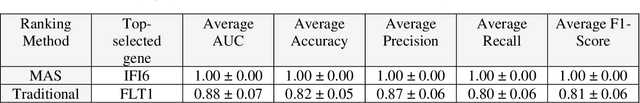

This study introduces the Supervised Magnitude-Altitude Scoring (SMAS) methodology, a machine learning-based approach, for analyzing gene expression data obtained from nonhuman primates (NHPs) infected with Ebola virus (EBOV). We utilize a comprehensive dataset of NanoString gene expression profiles from Ebola-infected NHPs, deploying the SMAS system for nuanced host-pathogen interaction analysis. SMAS effectively combines gene selection based on statistical significance and expression changes, employing linear classifiers such as logistic regression to accurately differentiate between RT-qPCR positive and negative NHP samples. A key finding of our research is the identification of IFI6 and IFI27 as critical biomarkers, demonstrating exceptional predictive performance with 100% accuracy and Area Under the Curve (AUC) metrics in classifying various stages of Ebola infection. Alongside IFI6 and IFI27, genes, including MX1, OAS1, and ISG15, were significantly upregulated, highlighting their essential roles in the immune response to EBOV. Our results underscore the efficacy of the SMAS method in revealing complex genetic interactions and response mechanisms during EBOV infection. This research provides valuable insights into EBOV pathogenesis and aids in developing more precise diagnostic tools and therapeutic strategies to address EBOV infection in particular and viral infection in general.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge