ivadomed: A Medical Imaging Deep Learning Toolbox
Paper and Code
Oct 20, 2020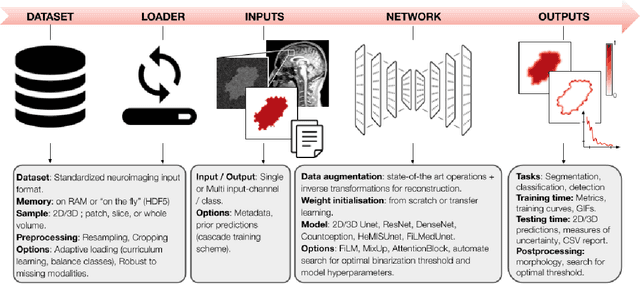
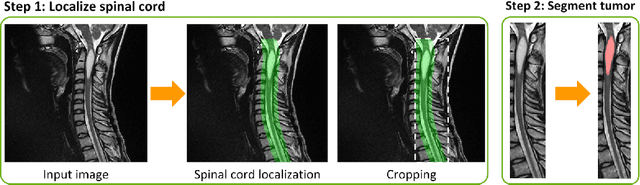
ivadomed is an open-source Python package for designing, end-to-end training, and evaluating deep learning models applied to medical imaging data. The package includes APIs, command-line tools, documentation, and tutorials. ivadomed also includes pre-trained models such as spinal tumor segmentation and vertebral labeling. Original features of ivadomed include a data loader that can parse image metadata (e.g., acquisition parameters, image contrast, resolution) and subject metadata (e.g., pathology, age, sex) for custom data splitting or extra information during training and evaluation. Any dataset following the Brain Imaging Data Structure (BIDS) convention will be compatible with ivadomed without the need to manually organize the data, which is typically a tedious task. Beyond the traditional deep learning methods, ivadomed features cutting-edge architectures, such as FiLM and HeMis, as well as various uncertainty estimation methods (aleatoric and epistemic), and losses adapted to imbalanced classes and non-binary predictions. Each step is conveniently configurable via a single file. At the same time, the code is highly modular to allow addition/modification of an architecture or pre/post-processing steps. Example applications of ivadomed include MRI object detection, segmentation, and labeling of anatomical and pathological structures. Overall, ivadomed enables easy and quick exploration of the latest advances in deep learning for medical imaging applications. ivadomed's main project page is available at https://ivadomed.org.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge