Inference for High Dimensional Censored Quantile Regression
Paper and Code
Jul 22, 2021
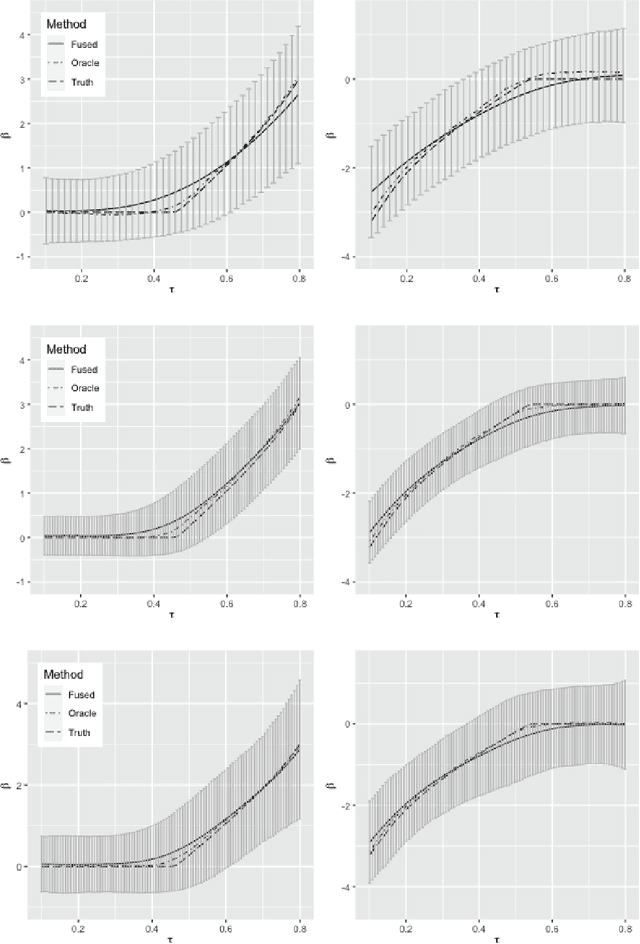

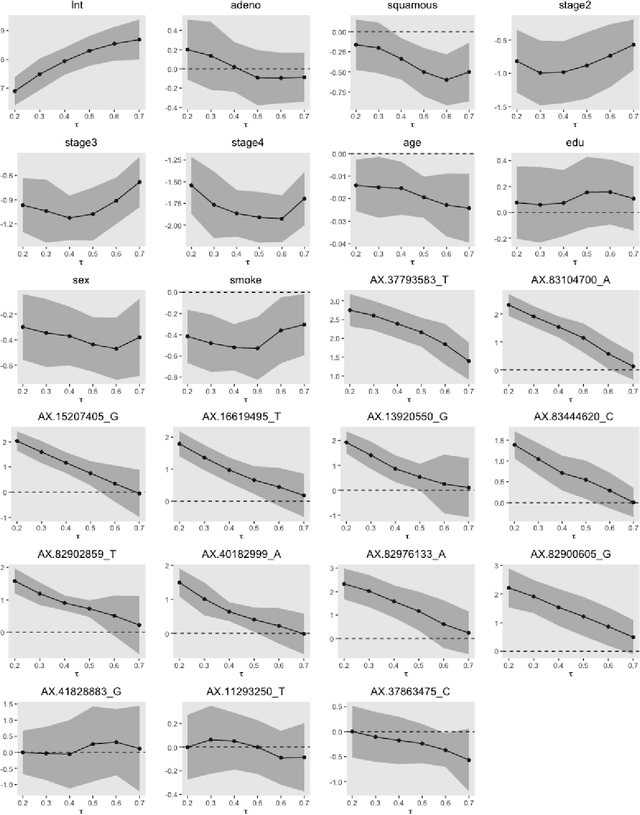
With the availability of high dimensional genetic biomarkers, it is of interest to identify heterogeneous effects of these predictors on patients' survival, along with proper statistical inference. Censored quantile regression has emerged as a powerful tool for detecting heterogeneous effects of covariates on survival outcomes. To our knowledge, there is little work available to draw inference on the effects of high dimensional predictors for censored quantile regression. This paper proposes a novel procedure to draw inference on all predictors within the framework of global censored quantile regression, which investigates covariate-response associations over an interval of quantile levels, instead of a few discrete values. The proposed estimator combines a sequence of low dimensional model estimates that are based on multi-sample splittings and variable selection. We show that, under some regularity conditions, the estimator is consistent and asymptotically follows a Gaussian process indexed by the quantile level. Simulation studies indicate that our procedure can properly quantify the uncertainty of the estimates in high dimensional settings. We apply our method to analyze the heterogeneous effects of SNPs residing in lung cancer pathways on patients' survival, using the Boston Lung Cancer Survival Cohort, a cancer epidemiology study on the molecular mechanism of lung cancer.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge