Harnessing spatial MRI normalization: patch individual filter layers for CNNs
Paper and Code
Nov 14, 2019
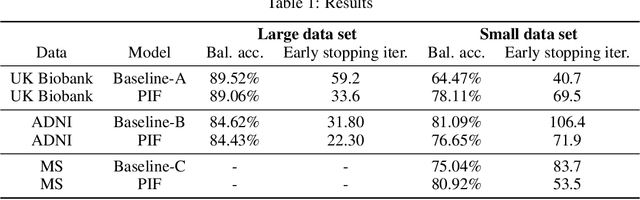
Neuroimaging studies based on magnetic resonance imaging (MRI) typically employ rigorous forms of preprocessing. Images are spatially normalized to a standard template using linear and non-linear transformations. Thus, one can assume that a patch at location (x, y, height, width) contains the same brain region across the entire data set. Most analyses applied on brain MRI using convolutional neural networks (CNNs) ignore this distinction from natural images. Here, we suggest a new layer type called patch individual filter (PIF) layer, which trains higher-level filters locally as we assume that more abstract features are locally specific after spatial normalization. We evaluate PIF layers on three different tasks, namely sex classification as well as either Alzheimer's disease (AD) or multiple sclerosis (MS) detection. We demonstrate that CNNs using PIF layers outperform their counterparts in several, especially low sample size settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge