Graph Autoencoders for Embedding Learning in Brain Networks and Major Depressive Disorder Identification
Paper and Code
Jul 27, 2021
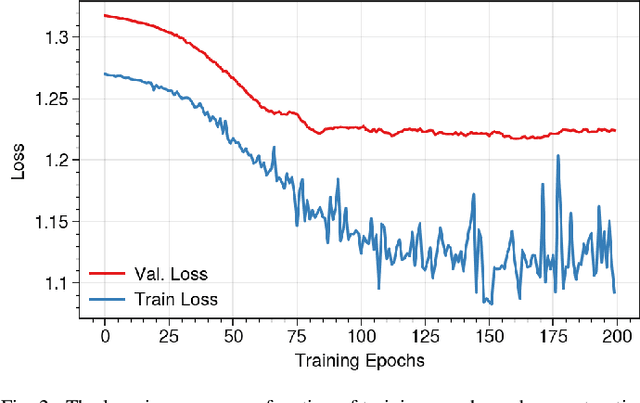
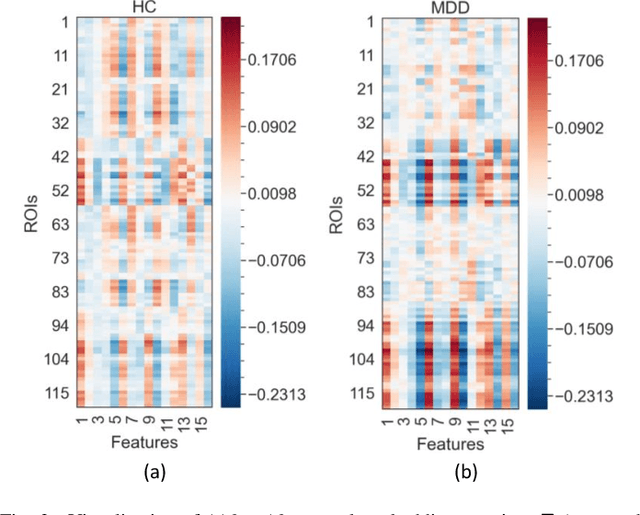
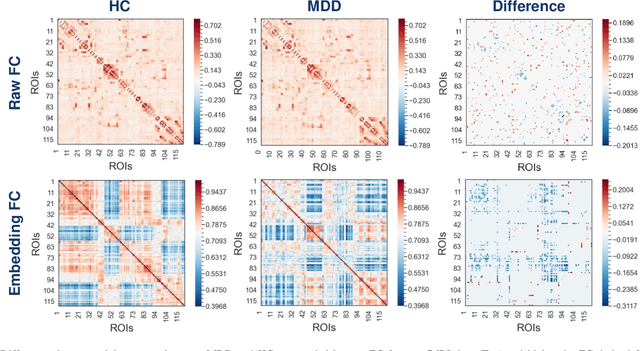
Brain functional connectivity (FC) reveals biomarkers for identification of various neuropsychiatric disorders. Recent application of deep neural networks (DNNs) to connectome-based classification mostly relies on traditional convolutional neural networks using input connectivity matrices on a regular Euclidean grid. We propose a graph deep learning framework to incorporate the non-Euclidean information about graph structure for classifying functional magnetic resonance imaging (fMRI)- derived brain networks in major depressive disorder (MDD). We design a novel graph autoencoder (GAE) architecture based on the graph convolutional networks (GCNs) to embed the topological structure and node content of large-sized fMRI networks into low-dimensional latent representations. In network construction, we employ the Ledoit-Wolf (LDW) shrinkage method to estimate the high-dimensional FC metrics efficiently from fMRI data. We consider both supervised and unsupervised approaches for the graph embedded learning. The learned embeddings are then used as feature inputs for a deep fully-connected neural network (FCNN) to discriminate MDD from healthy controls. Evaluated on a resting-state fMRI MDD dataset with 43 subjects, results show that the proposed GAE-FCNN model significantly outperforms several state-of-the-art DNN methods for brain connectome classification, achieving accuracy of 72.50% using the LDW-FC metrics as node features. The graph embeddings of fMRI FC networks learned by the GAE also reveal apparent group differences between MDD and HC. Our new framework demonstrates feasibility of learning graph embeddings on brain networks to provide discriminative information for diagnosis of brain disorders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge