Grad-CAMO: Learning Interpretable Single-Cell Morphological Profiles from 3D Cell Painting Images
Paper and Code
Mar 26, 2024
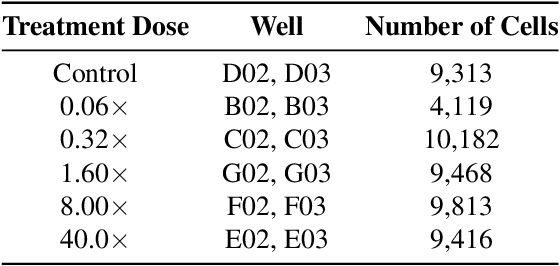
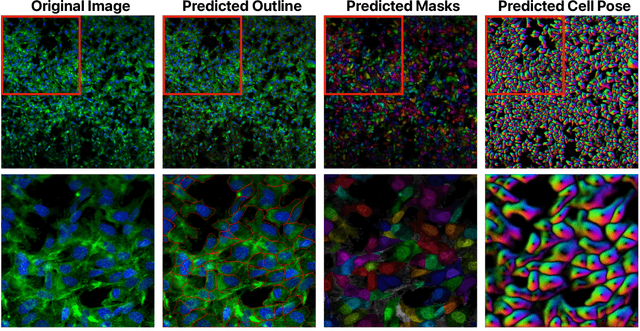
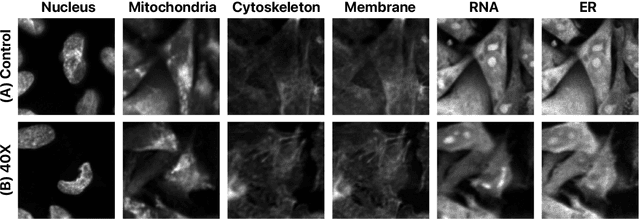
Despite their black-box nature, deep learning models are extensively used in image-based drug discovery to extract feature vectors from single cells in microscopy images. To better understand how these networks perform representation learning, we employ visual explainability techniques (e.g., Grad-CAM). Our analyses reveal several mechanisms by which supervised models cheat, exploiting biologically irrelevant pixels when extracting morphological features from images, such as noise in the background. This raises doubts regarding the fidelity of learned single-cell representations and their relevance when investigating downstream biological questions. To address this misalignment between researcher expectations and machine behavior, we introduce Grad-CAMO, a novel single-cell interpretability score for supervised feature extractors. Grad-CAMO measures the proportion of a model's attention that is concentrated on the cell of interest versus the background. This metric can be assessed per-cell or averaged across a validation set, offering a tool to audit individual features vectors or guide the improved design of deep learning architectures. Importantly, Grad-CAMO seamlessly integrates into existing workflows, requiring no dataset or model modifications, and is compatible with both 2D and 3D Cell Painting data. Additional results are available at https://github.com/eigenvivek/Grad-CAMO.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge