Gigapixel Whole-Slide Images Classification using Locally Supervised Learning
Paper and Code
Jul 17, 2022
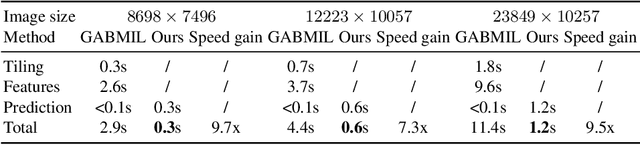
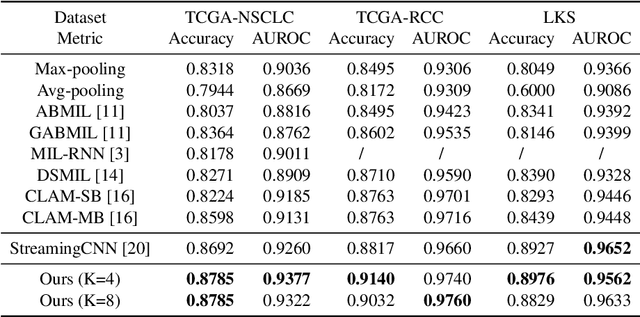


Histopathology whole slide images (WSIs) play a very important role in clinical studies and serve as the gold standard for many cancer diagnoses. However, generating automatic tools for processing WSIs is challenging due to their enormous sizes. Currently, to deal with this issue, conventional methods rely on a multiple instance learning (MIL) strategy to process a WSI at patch level. Although effective, such methods are computationally expensive, because tiling a WSI into patches takes time and does not explore the spatial relations between these tiles. To tackle these limitations, we propose a locally supervised learning framework which processes the entire slide by exploring the entire local and global information that it contains. This framework divides a pre-trained network into several modules and optimizes each module locally using an auxiliary model. We also introduce a random feature reconstruction unit (RFR) to preserve distinguishing features during training and improve the performance of our method by 1% to 3%. Extensive experiments on three publicly available WSI datasets: TCGA-NSCLC, TCGA-RCC and LKS, highlight the superiority of our method on different classification tasks. Our method outperforms the state-of-the-art MIL methods by 2% to 5% in accuracy, while being 7 to 10 times faster. Additionally, when dividing it into eight modules, our method requires as little as 20% of the total gpu memory required by end-to-end training. Our code is available at https://github.com/cvlab-stonybrook/local_learning_wsi.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge