DProQ: A Gated-Graph Transformer for Protein Complex Structure Assessment
Paper and Code
May 21, 2022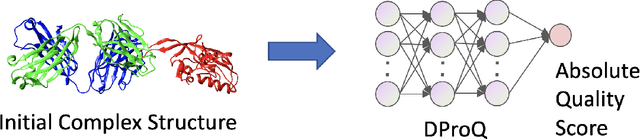



Proteins interact to form complexes to carry out essential biological functions. Computational methods have been developed to predict the structures of protein complexes. However, an important challenge in protein complex structure prediction is to estimate the quality of predicted protein complex structures without any knowledge of the corresponding native structures. Such estimations can then be used to select high-quality predicted complex structures to facilitate biomedical research such as protein function analysis and drug discovery. We challenge this significant task with DProQ, which introduces a gated neighborhood-modulating Graph Transformer (GGT) designed to predict the quality of 3D protein complex structures. Notably, we incorporate node and edge gates within a novel Graph Transformer framework to control information flow during graph message passing. We train and evaluate DProQ on four newly-developed datasets that we make publicly available in this work. Our rigorous experiments demonstrate that DProQ achieves state-of-the-art performance in ranking protein complex structures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge